Overview
51/51
Variable Coverage
All ACE atmosphere variables found
1/2
Checks Passed
1 components with issues
1.34x
Storage Ratio
FME 2.11 GB vs Legacy 1.57 GB
N/A
Timing Overhead
No timing data available
Component Summary
| Component | Status | Issues |
|---|---|---|
| EAM | PASS | 0 |
| Cross-Verify | FAIL | 6 |
Show all 6 issues
| Component | Issue |
|---|---|
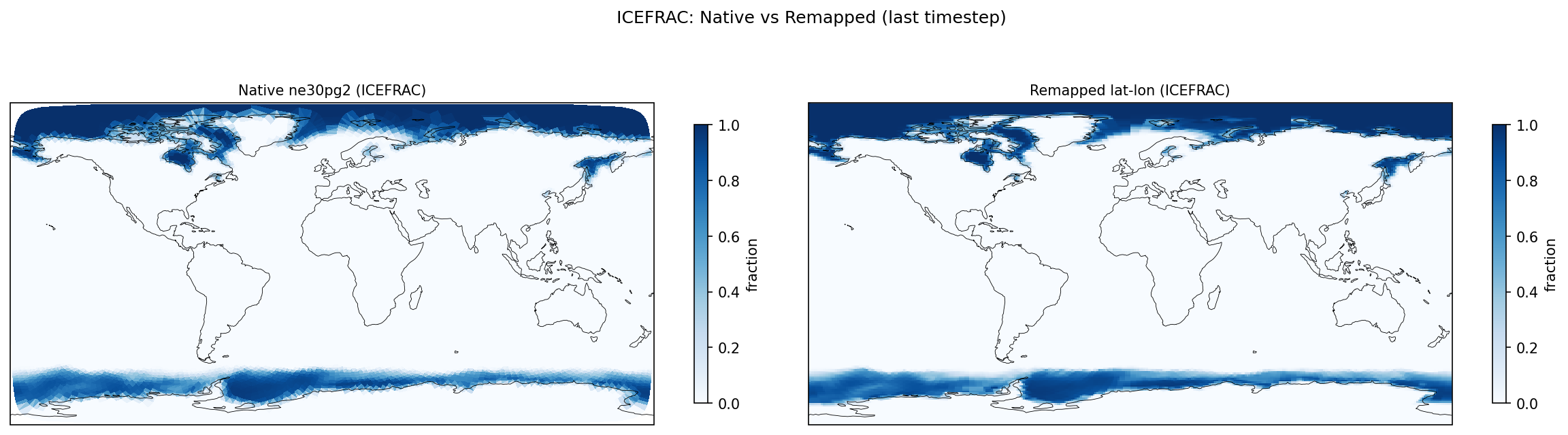
| Cross-Verify | GMEAN ICEFRAC: rel_diff = 0.0966 |
| Cross-Verify | GMEAN FSUS: rel_diff = 0.05457 |
| Cross-Verify | GMEAN TAUX: rel_diff = 0.3693 |
| Cross-Verify | GMEAN TAUY: rel_diff = 0.02138 |
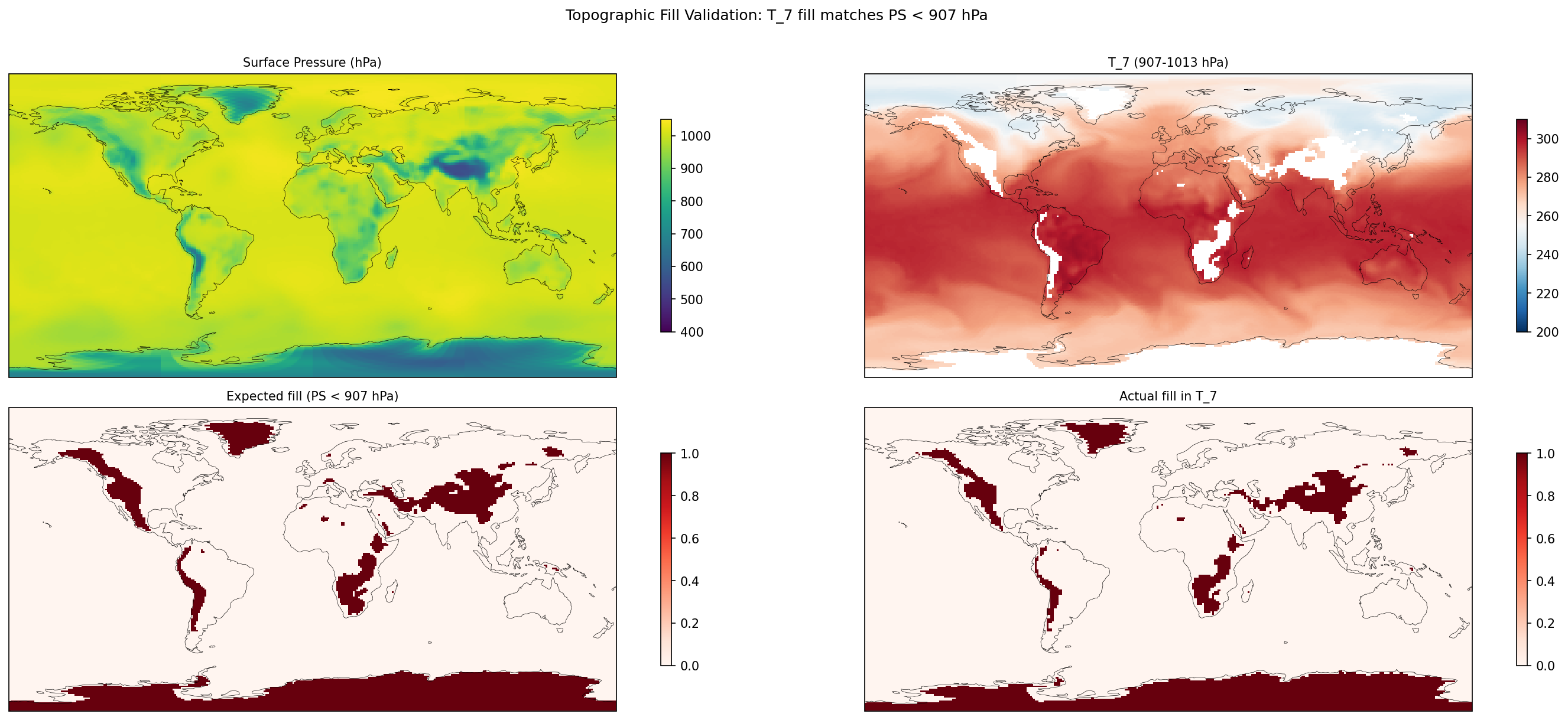
| Cross-Verify | TOPO_FILL: max fill mismatch 1.590% |
| Cross-Verify | LINEARITY STW: max|rel| = 2.46e-01 |
ACE Atmosphere Variable Requirements
51/51 variables found all present
YAML Variable Specification (ACE canonical names)
## ATMOSPHERE next_step_forcing_names: - SOLIN in_names: - land_fraction - ocean_fraction - sea_ice_fraction - PHIS - SOLIN - PS - TS - T_0 - T_1 - T_2 - T_3 - T_4 - T_5 - T_6 - T_7 - specific_total_water_0 - specific_total_water_1 - specific_total_water_2 - specific_total_water_3 - specific_total_water_4 - specific_total_water_5 - specific_total_water_6 - specific_total_water_7 - U_0 - U_1 - U_2 - U_3 - U_4 - U_5 - U_6 - U_7 - V_0 - V_1 - V_2 - V_3 - V_4 - V_5 - V_6 - V_7 out_names: - PS - TS - T_0 - T_1 - T_2 - T_3 - T_4 - T_5 - T_6 - T_7 - specific_total_water_0 - specific_total_water_1 - specific_total_water_2 - specific_total_water_3 - specific_total_water_4 - specific_total_water_5 - specific_total_water_6 - specific_total_water_7 - U_0 - U_1 - U_2 - U_3 - U_4 - U_5 - U_6 - U_7 - V_0 - V_1 - V_2 - V_3 - V_4 - V_5 - V_6 - V_7 - LHFLX - SHFLX - surface_precipitation_rate - surface_upward_longwave_flux - FLUT - FLDS - FSDS - surface_upward_shortwave_flux - top_of_atmos_upward_shortwave_flux - tendency_of_total_water_path_due_to_advection - TAUX - TAUY
Variable Coverage
| ACE Name | EAM Name | Category | Status |
|---|---|---|---|
SOLIN | SOLIN | forcing, in | FOUND |
land_fraction | LANDFRAC | in | FOUND |
ocean_fraction | OCNFRAC | in | FOUND |
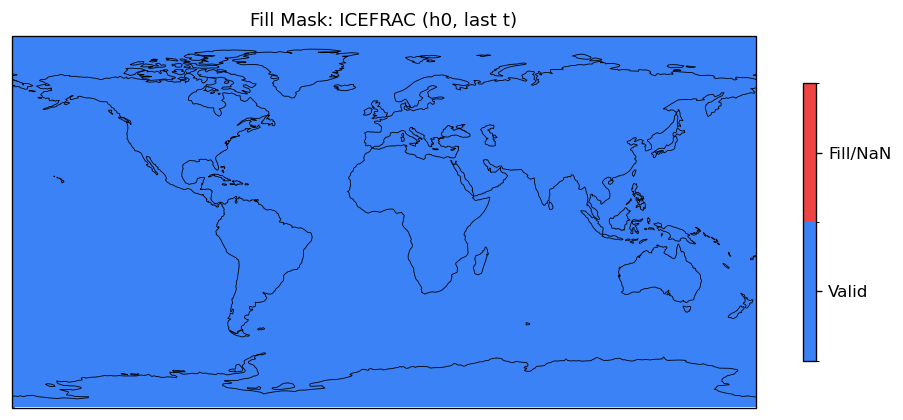
sea_ice_fraction | ICEFRAC | in | FOUND |
PHIS | PHIS | in | FOUND |
PS | PS | in, out | FOUND |
TS | TS | in, out | FOUND |
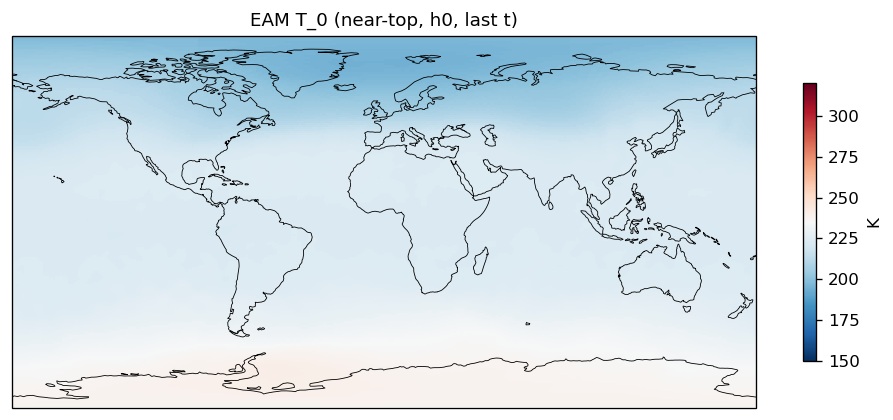
T_0 | T_0 | in, out | FOUND |
T_1 | T_1 | in, out | FOUND |
T_2 | T_2 | in, out | FOUND |
T_3 | T_3 | in, out | FOUND |
T_4 | T_4 | in, out | FOUND |
T_5 | T_5 | in, out | FOUND |
T_6 | T_6 | in, out | FOUND |
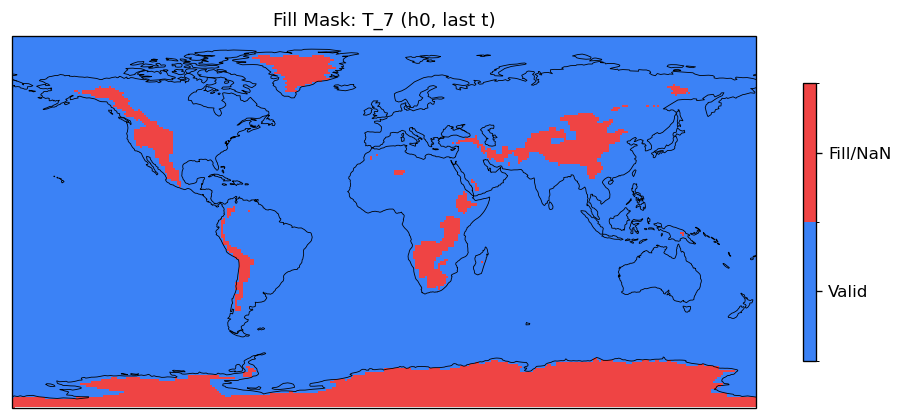
T_7 | T_7 | in, out | FOUND |
specific_total_water_0 | STW_0 | in, out | FOUND |
specific_total_water_1 | STW_1 | in, out | FOUND |
specific_total_water_2 | STW_2 | in, out | FOUND |
specific_total_water_3 | STW_3 | in, out | FOUND |
specific_total_water_4 | STW_4 | in, out | FOUND |
specific_total_water_5 | STW_5 | in, out | FOUND |
specific_total_water_6 | STW_6 | in, out | FOUND |
specific_total_water_7 | STW_7 | in, out | FOUND |
U_0 | U_0 | in, out | FOUND |
U_1 | U_1 | in, out | FOUND |
U_2 | U_2 | in, out | FOUND |
U_3 | U_3 | in, out | FOUND |
U_4 | U_4 | in, out | FOUND |
U_5 | U_5 | in, out | FOUND |
U_6 | U_6 | in, out | FOUND |
U_7 | U_7 | in, out | FOUND |
V_0 | V_0 | in, out | FOUND |
V_1 | V_1 | in, out | FOUND |
V_2 | V_2 | in, out | FOUND |
V_3 | V_3 | in, out | FOUND |
V_4 | V_4 | in, out | FOUND |
V_5 | V_5 | in, out | FOUND |
V_6 | V_6 | in, out | FOUND |
V_7 | V_7 | in, out | FOUND |
LHFLX | LHFLX | out | FOUND |
SHFLX | SHFLX | out | FOUND |
surface_precipitation_rate | PRECT | out | FOUND |
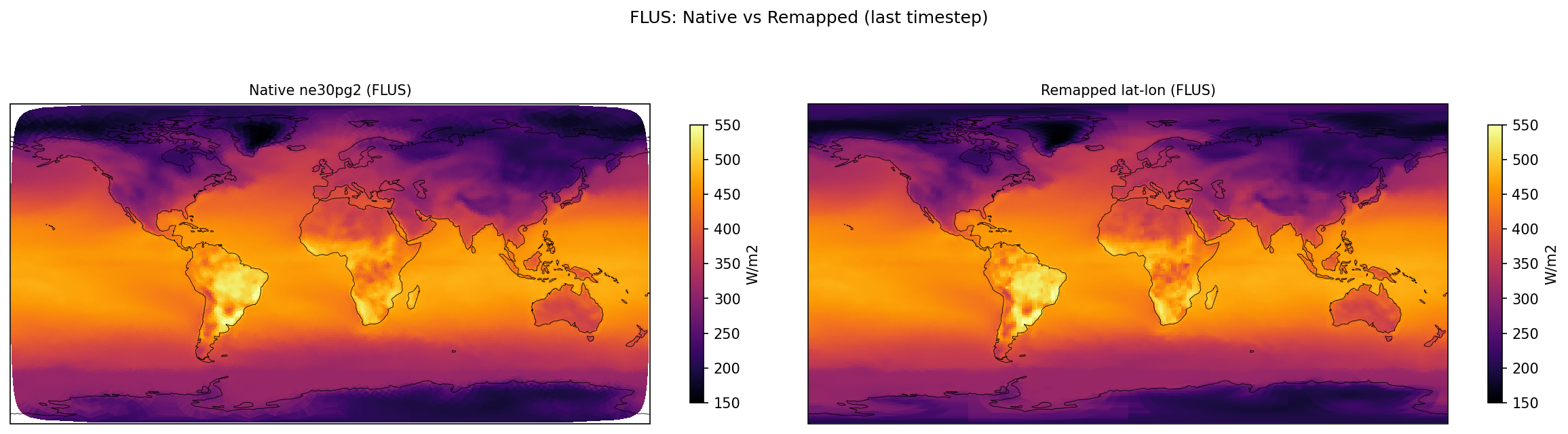
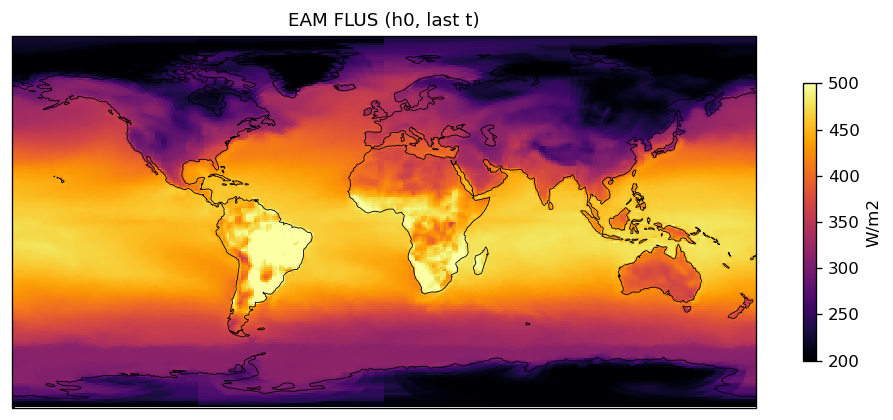
surface_upward_longwave_flux | FLUS | out | FOUND |
FLUT | FLUT | out | FOUND |
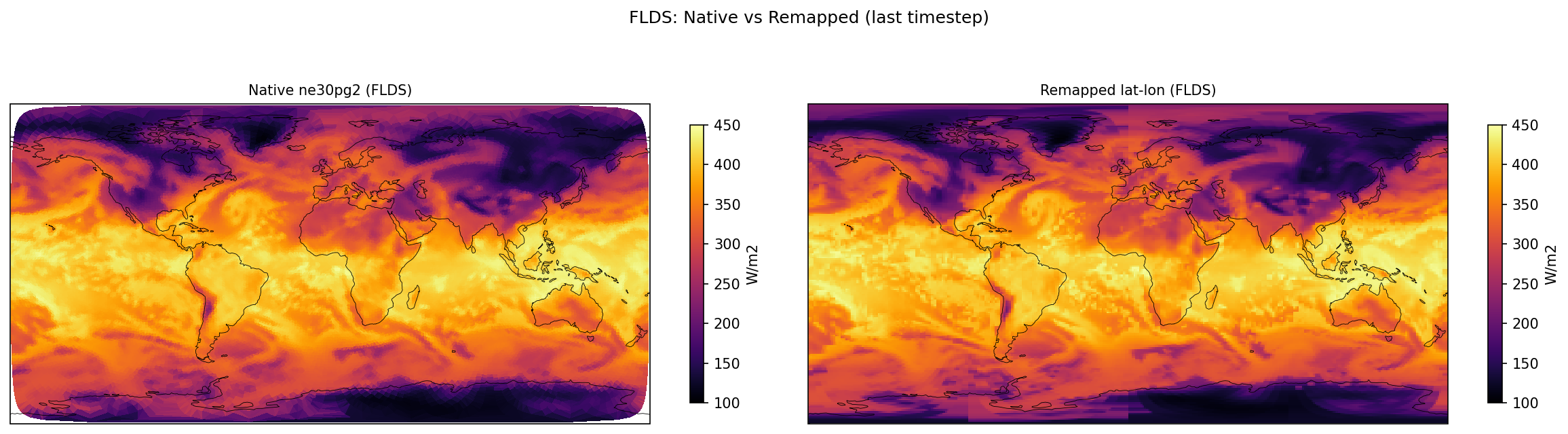
FLDS | FLDS | out | FOUND |
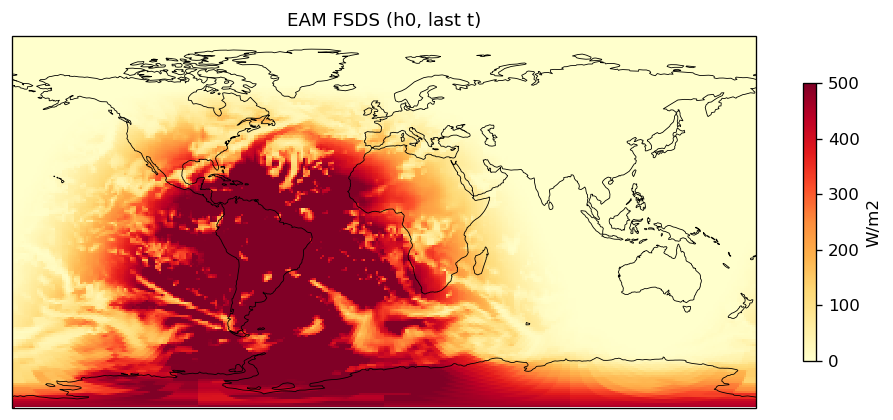
FSDS | FSDS | out | FOUND |
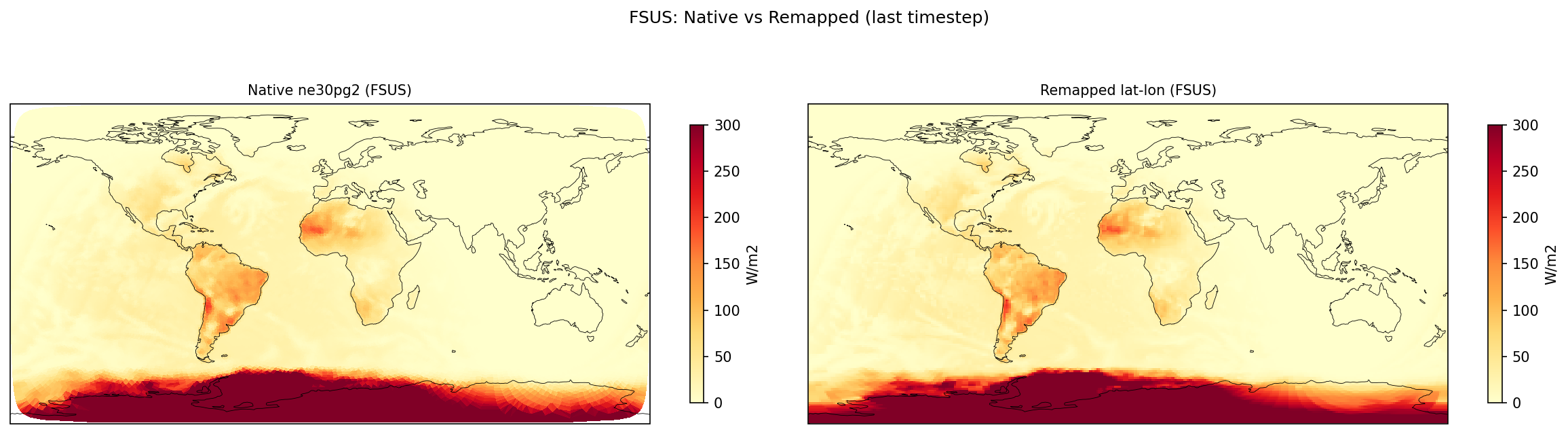
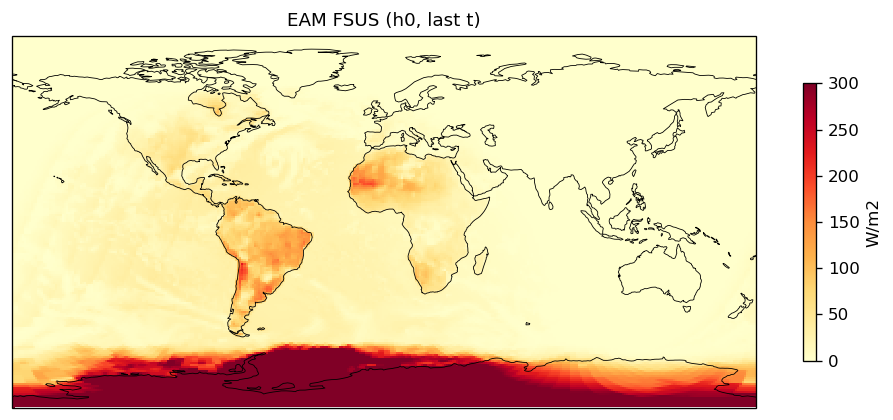
surface_upward_shortwave_flux | FSUS | out | FOUND |
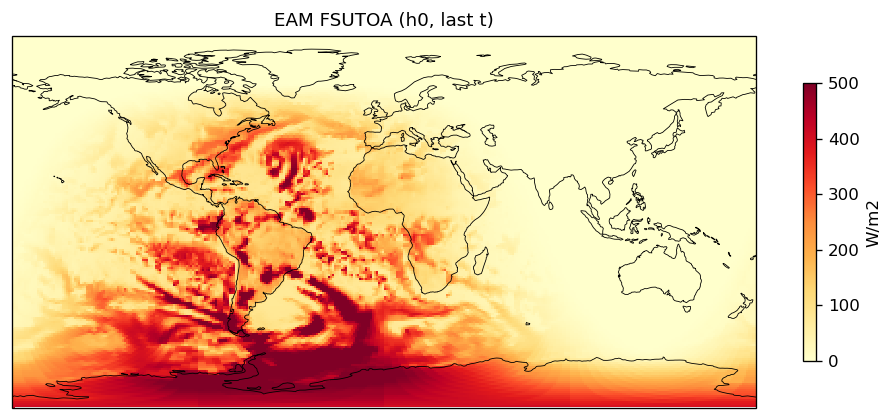
top_of_atmos_upward_shortwave_flux | FSUTOA | out | FOUND |
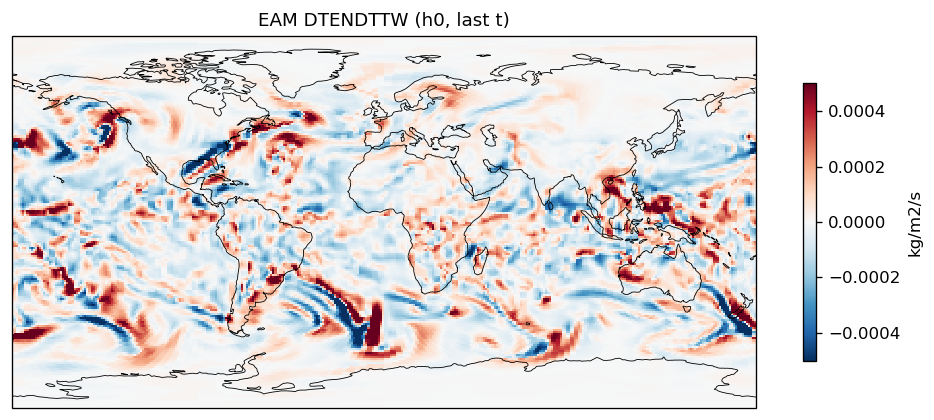
tendency_of_total_water_path_due_to_advection | DTENDTTW | out | FOUND |
TAUX | TAUX | out | FOUND |
TAUY | TAUY | out | FOUND |
File Inventory
| Category | File Count |
|---|---|
| EAM h0 | 3 |
Fill / NaN Report
Variable-level fill fraction details
| Component | Variable | Total Cells | Valid | Fill/NaN | Fill % |
|---|---|---|---|---|---|
| EAM | h0/PHIS | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/PS | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/TS | 259,200 | 259,200 | 0 | 0.0% |
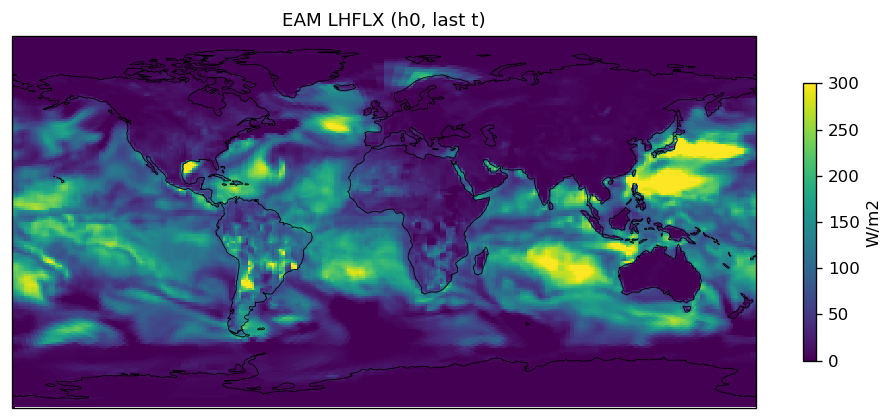
| EAM | h0/LHFLX | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/SHFLX | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/FLDS | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/FSDS | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/FSNTOA | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/FSUTOA | 259,200 | 259,200 | 0 | 0.0% |
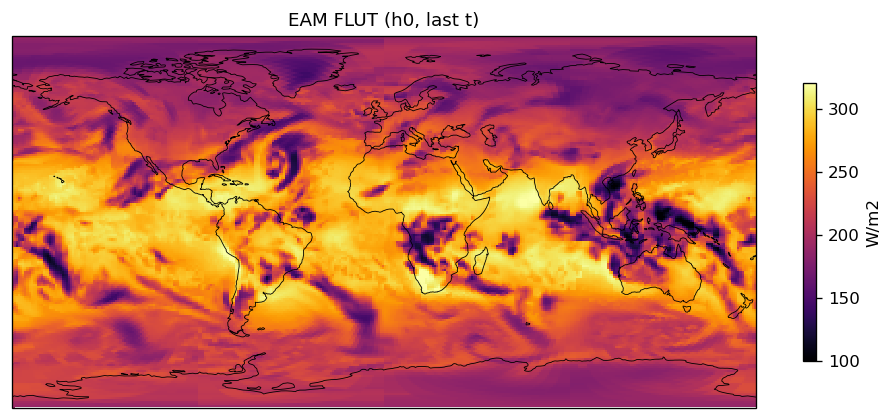
| EAM | h0/FLUT | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/SOLIN | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/FSUS | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/FLUS | 259,200 | 259,200 | 0 | 0.0% |
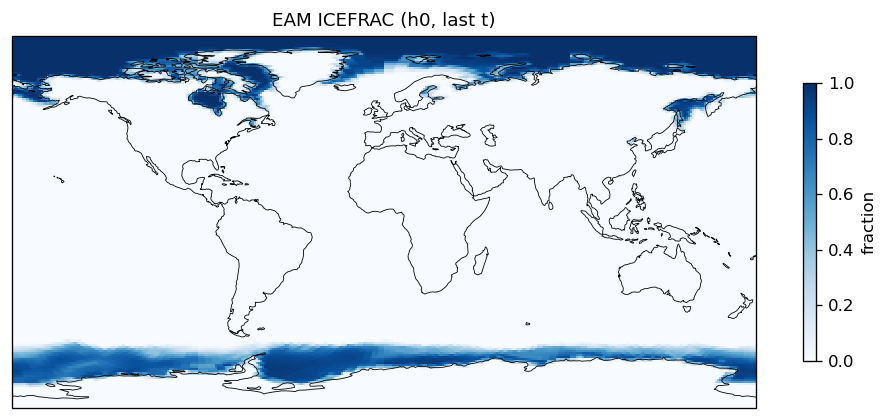
| EAM | h0/ICEFRAC | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/LANDFRAC | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/OCNFRAC | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/TAUX | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/TAUY | 259,200 | 259,200 | 0 | 0.0% |
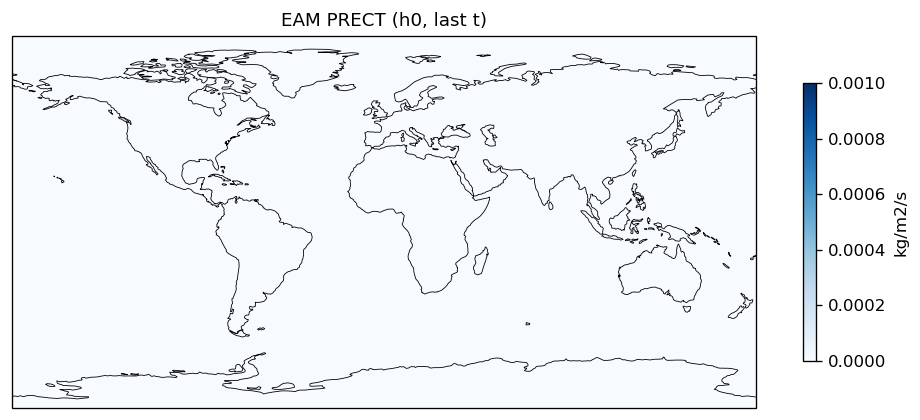
| EAM | h0/PRECT | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/DTENDTTW | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/T_0 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/T_1 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/T_2 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/T_3 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/T_4 | 259,200 | 259,200 | 0 | 0.0% |
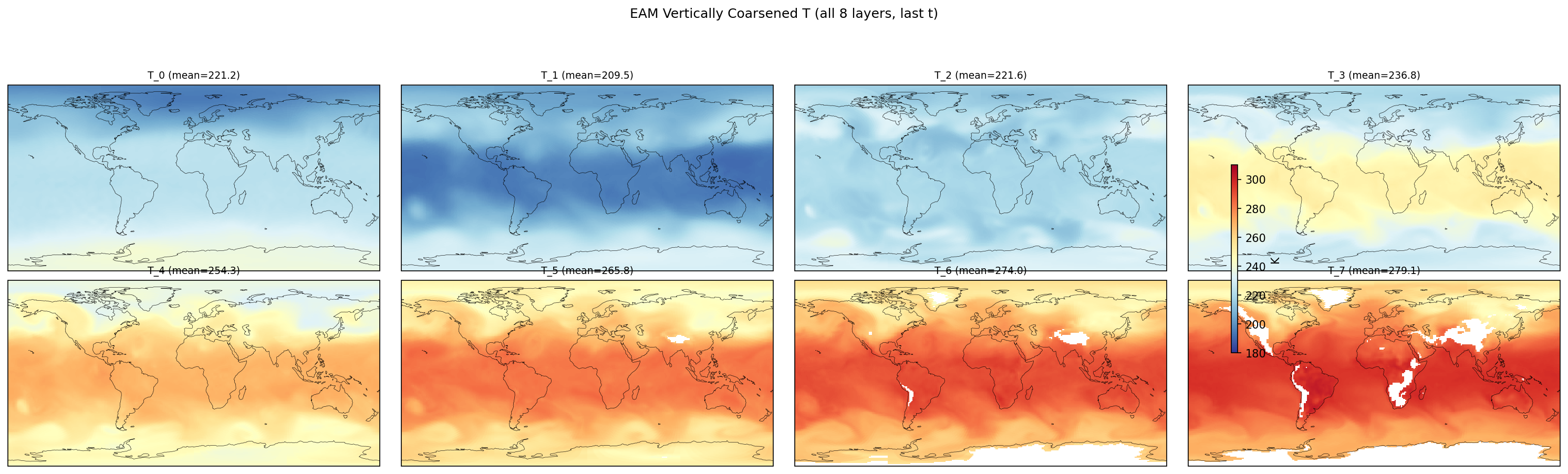
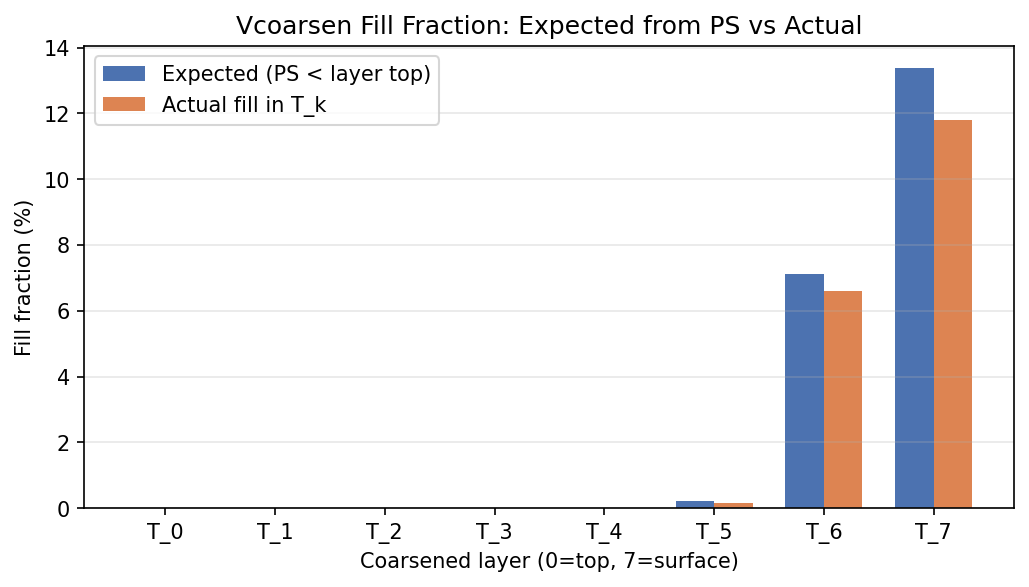
| EAM | h0/T_5 | 259,200 | 258,765 | 435 | 0.2% |
| EAM | h0/T_6 | 259,200 | 242,122 | 17,078 | 6.6% |
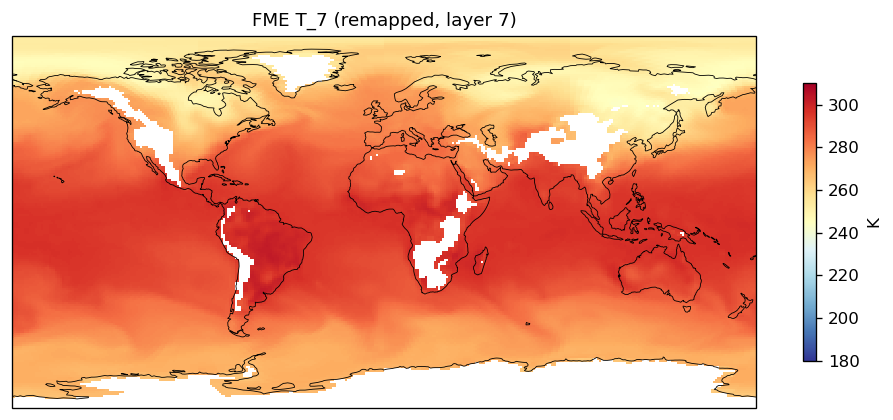
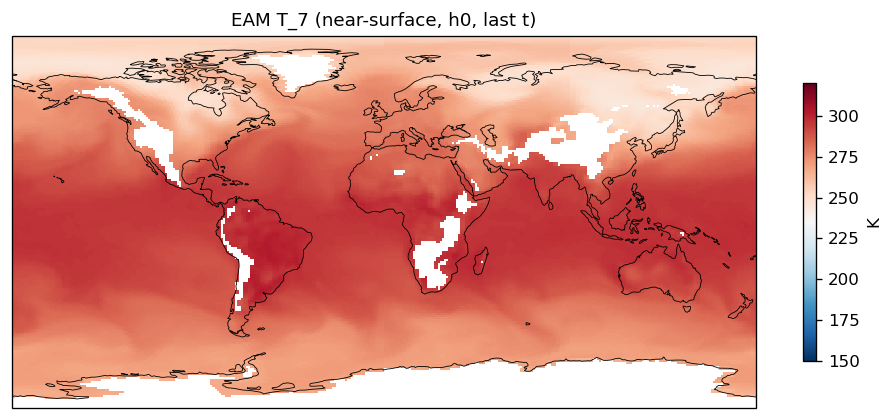
| EAM | h0/T_7 | 259,200 | 228,443 | 30,757 | 11.9% |
| EAM | h0/STW_0 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/STW_1 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/STW_2 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/STW_3 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/STW_4 | 259,200 | 259,200 | 0 | 0.0% |
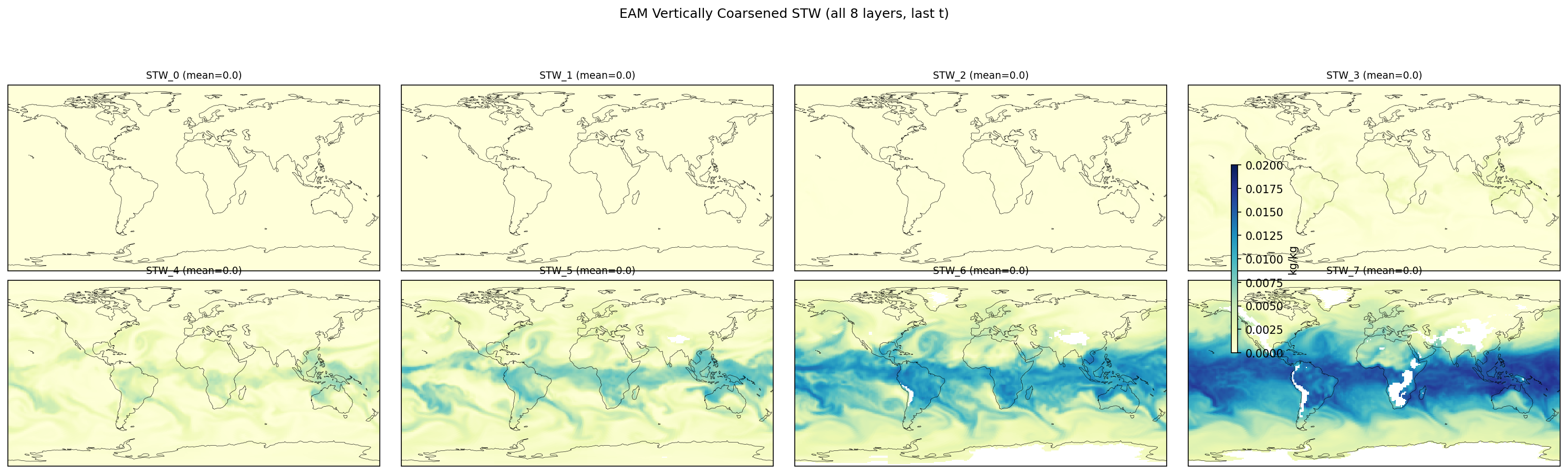
| EAM | h0/STW_5 | 259,200 | 258,765 | 435 | 0.2% |
| EAM | h0/STW_6 | 259,200 | 242,122 | 17,078 | 6.6% |
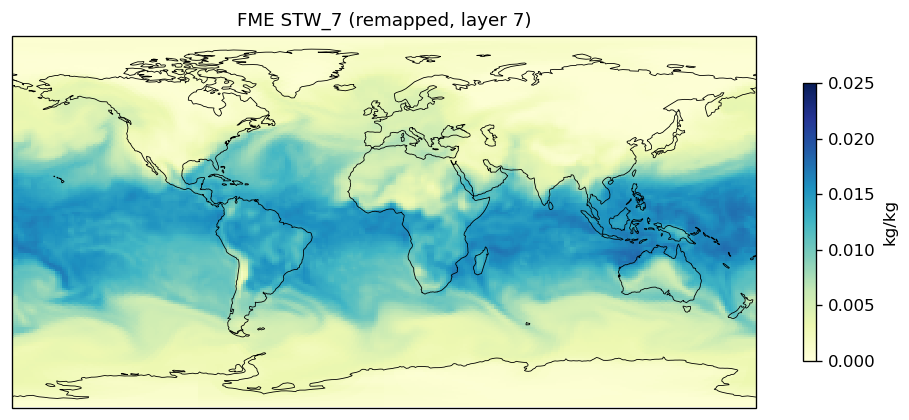
| EAM | h0/STW_7 | 259,200 | 228,443 | 30,757 | 11.9% |
| EAM | h0/U_0 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/U_1 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/U_2 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/U_3 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/U_4 | 259,200 | 259,200 | 0 | 0.0% |
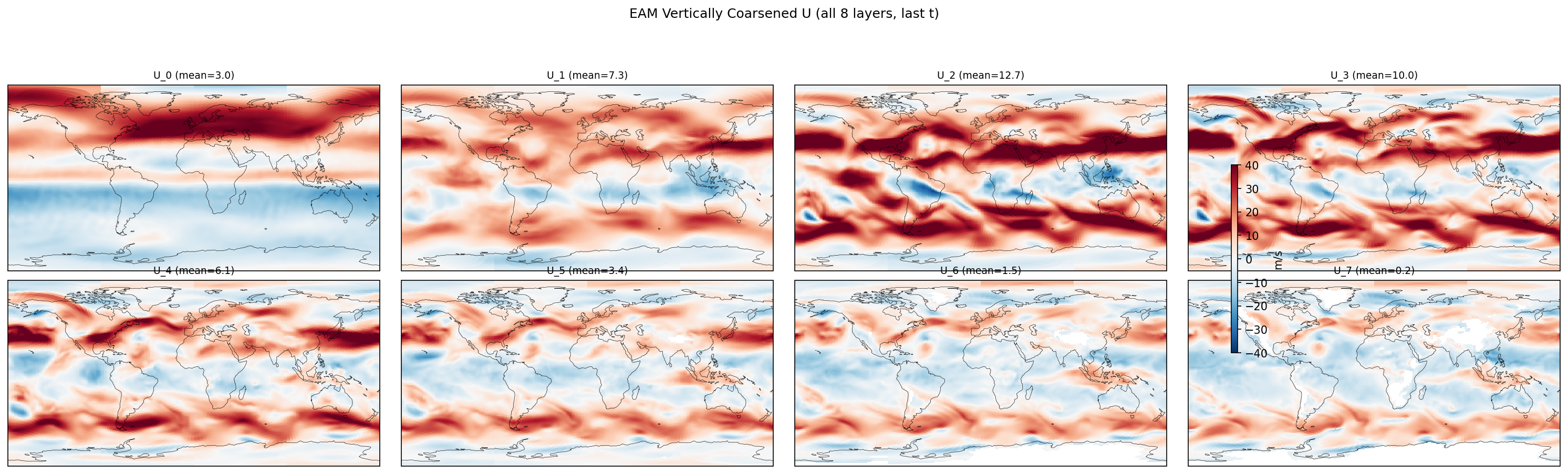
| EAM | h0/U_5 | 259,200 | 258,765 | 435 | 0.2% |
| EAM | h0/U_6 | 259,200 | 242,122 | 17,078 | 6.6% |
| EAM | h0/U_7 | 259,200 | 228,443 | 30,757 | 11.9% |
| EAM | h0/V_0 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/V_1 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/V_2 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/V_3 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/V_4 | 259,200 | 259,200 | 0 | 0.0% |
| EAM | h0/V_5 | 259,200 | 258,765 | 435 | 0.2% |
| EAM | h0/V_6 | 259,200 | 242,122 | 17,078 | 6.6% |
| EAM | h0/V_7 | 259,200 | 228,443 | 30,757 | 11.9% |
Cross-Verification: FME vs Legacy
Both cases run identical physics from the same ICs. FME diagnostics are output-only and do not modify model state. 22 pass, 6 fail, 1 info
Area-Weighted Global-Mean Comparison (cos-lat weighted, <2% tolerance) (18/22)
| Field | Status | Detail |
|---|---|---|
| PS | PASS | fme=98541.6, leg=98604.1, rel=6.34e-04 |
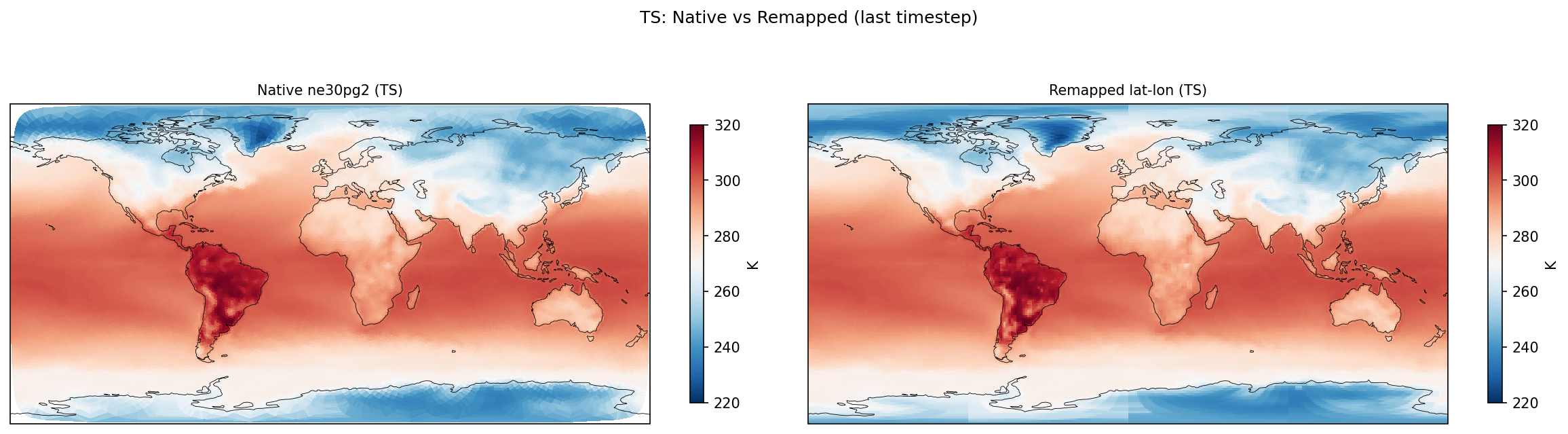
| TS | PASS | fme=286.599, leg=286.705, rel=3.71e-04 |
| PHIS | PASS | fme=2269.56, leg=2233.85, rel=1.60e-02 |
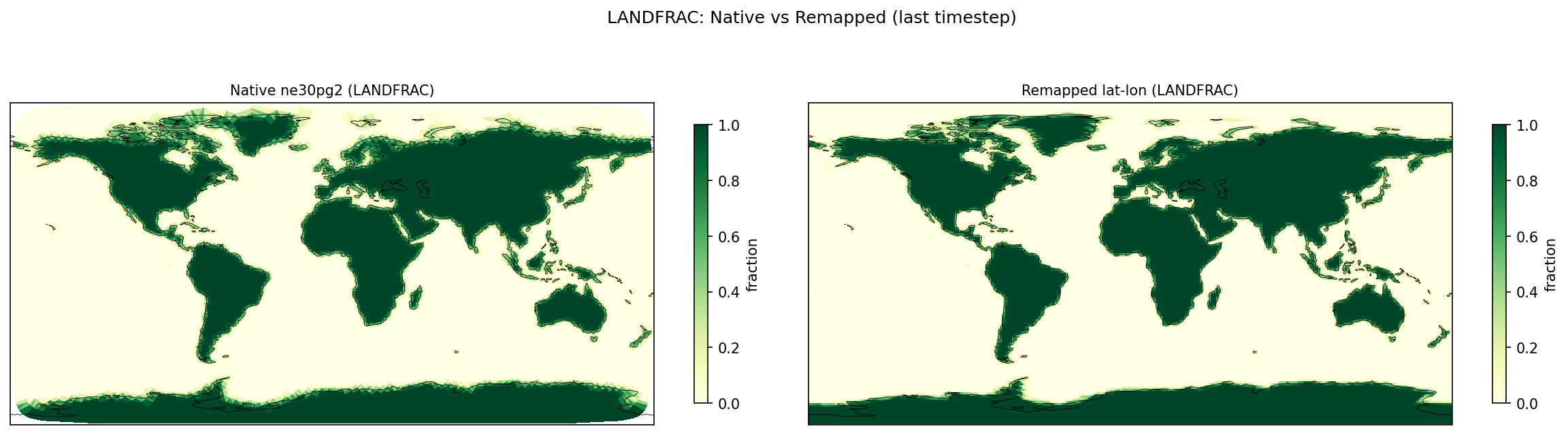
| LANDFRAC | PASS | fme=0.293414, leg=0.292714, rel=2.39e-03 |
| OCNFRAC | PASS | fme=0.649502, leg=0.655231, rel=8.74e-03 |
| ICEFRAC | DIFF | fme=0.0570837, leg=0.052055, rel=9.66e-02 |
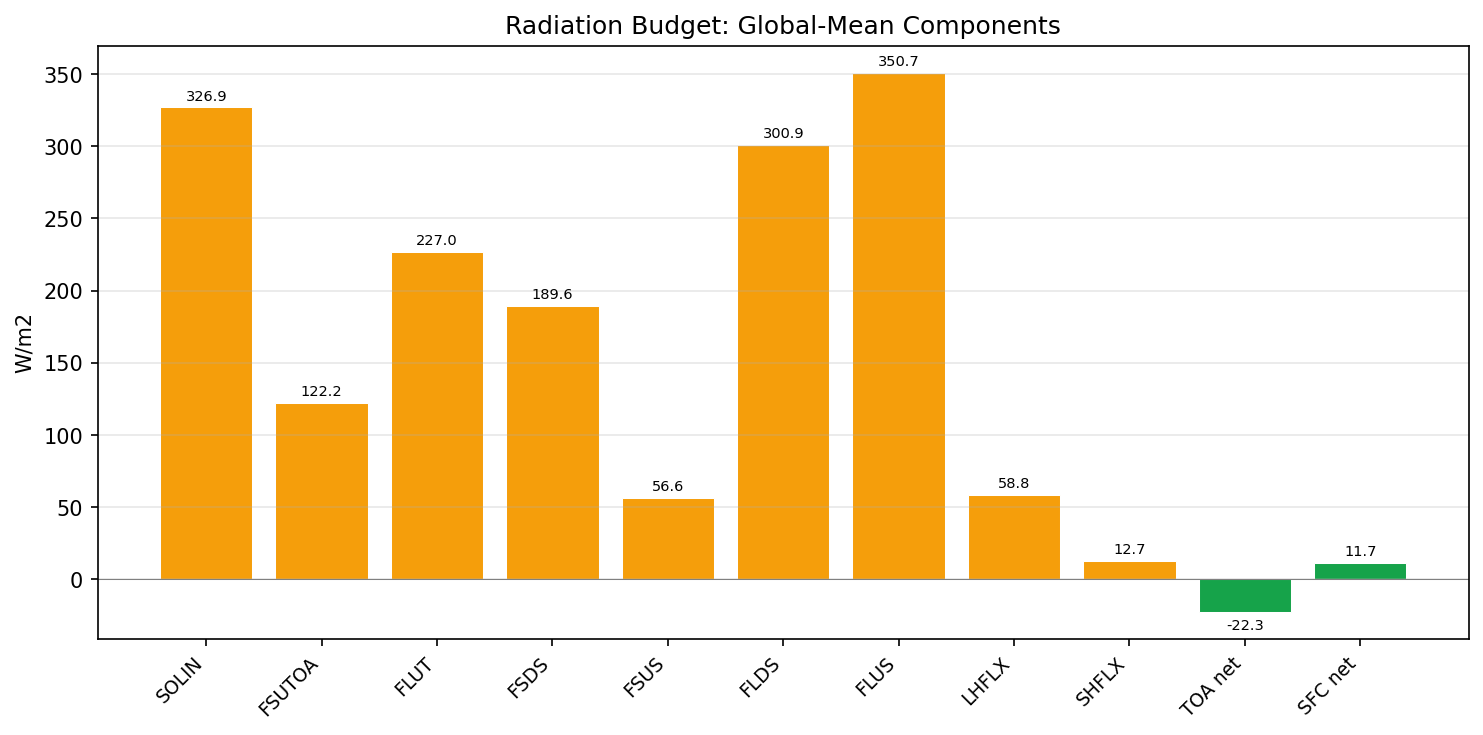
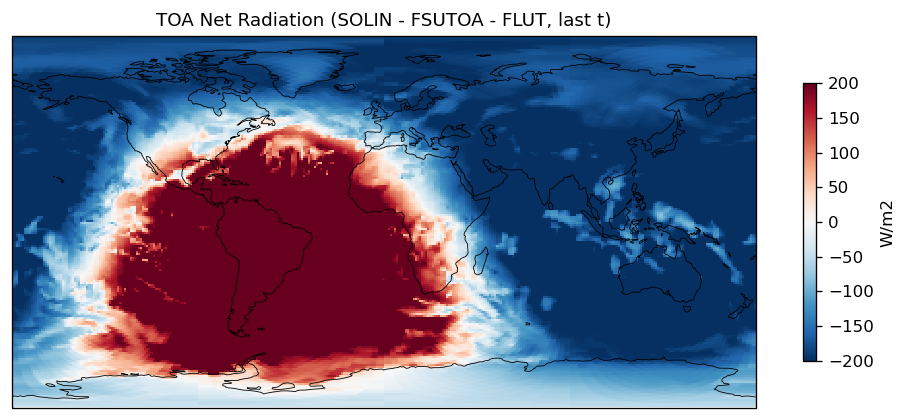
| SOLIN | PASS | fme=351.805, leg=351.571, rel=6.68e-04 |
| FSNTOA | PASS | fme=246.991, leg=247.372, rel=1.54e-03 |
| FSUTOA | PASS | fme=104.815, leg=104.199, rel=5.91e-03 |
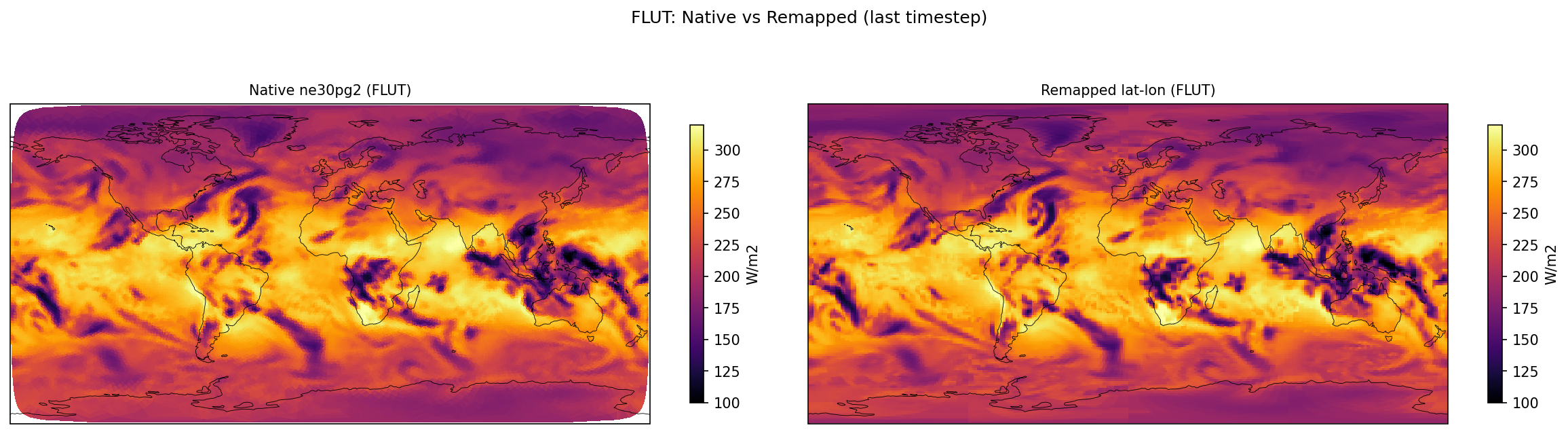
| FLUT | PASS | fme=239.598, leg=239.47, rel=5.37e-04 |
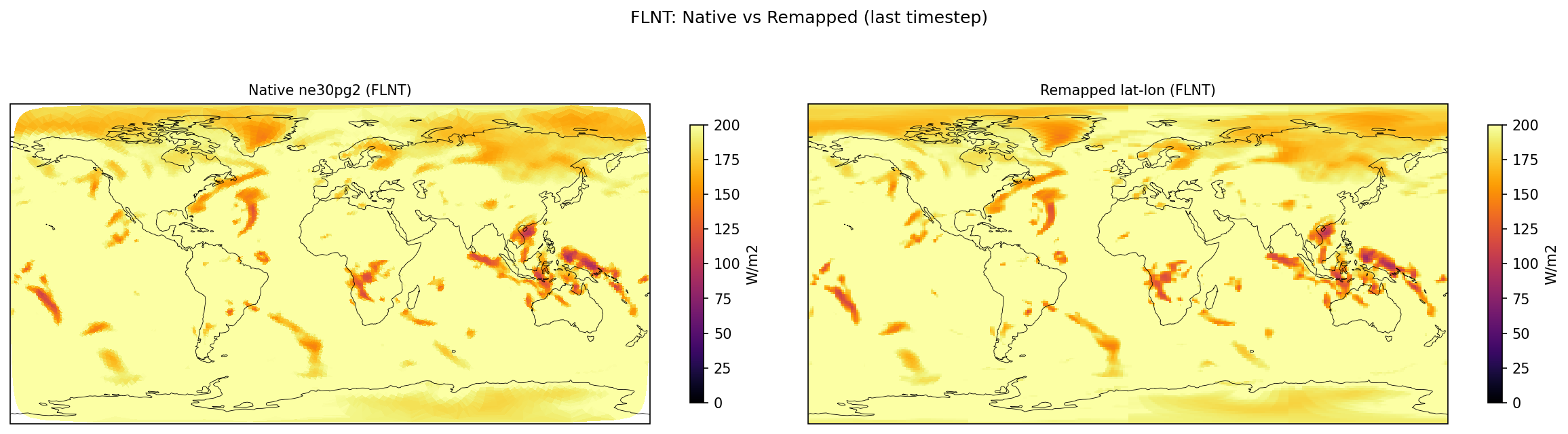
| FLNT | PASS | fme=239.4, leg=239.272, rel=5.36e-04 |
| FLDS | PASS | fme=337.223, leg=337.358, rel=4.01e-04 |
| FLUS | PASS | fme=391.271, leg=391.478, rel=5.29e-04 |
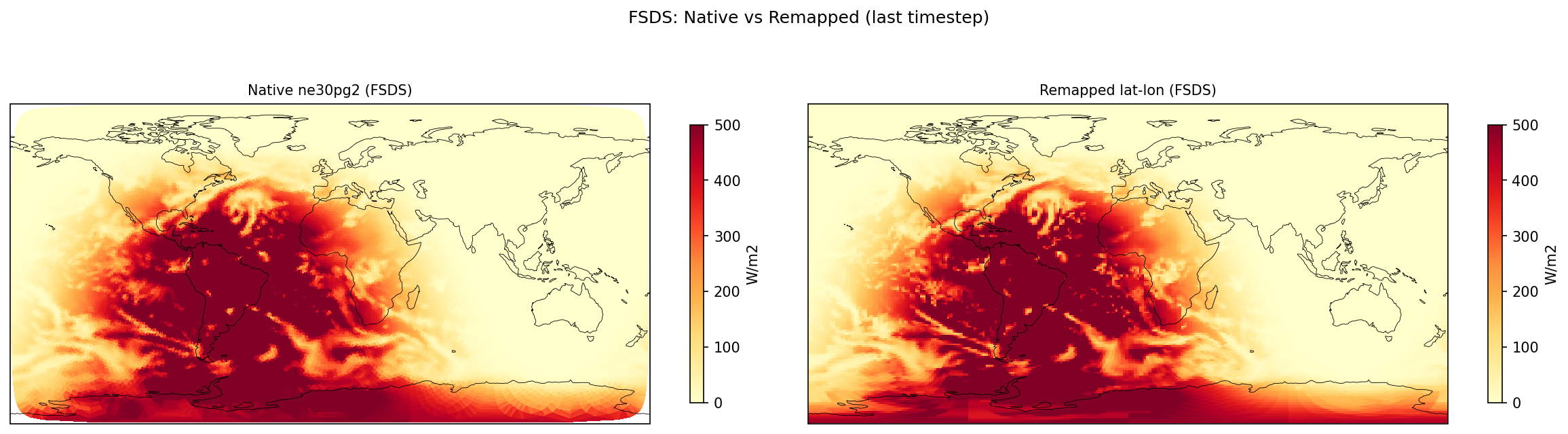
| FSDS | PASS | fme=197.113, leg=196.032, rel=5.52e-03 |
| FSUS | DIFF | fme=29.8047, leg=28.2625, rel=5.46e-02 |
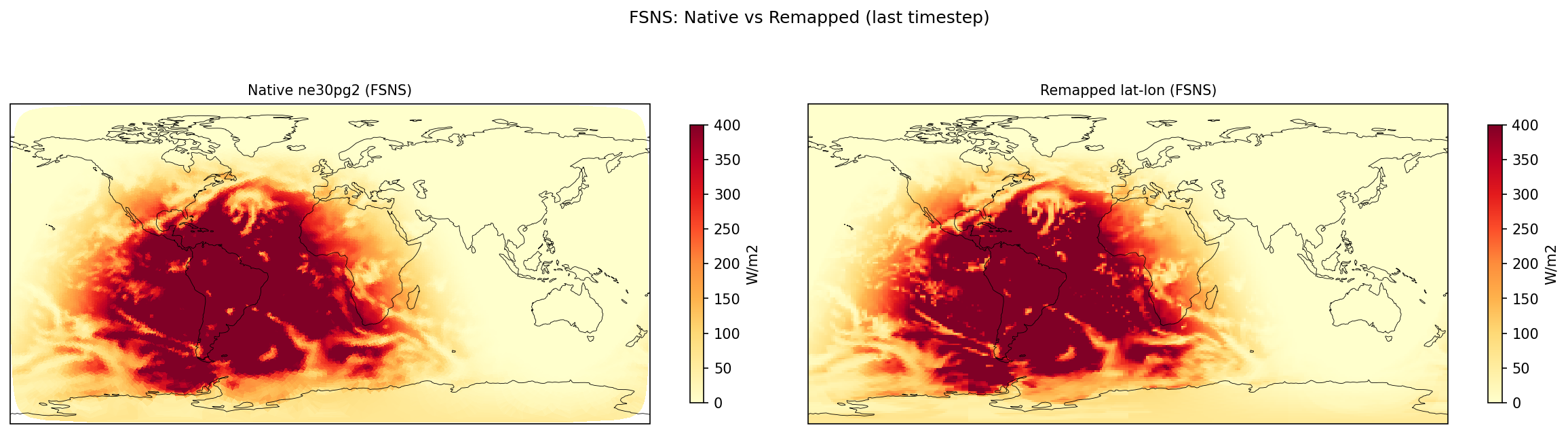
| FSNS | PASS | fme=167.309, leg=167.769, rel=2.74e-03 |
| FLNS | PASS | fme=54.0483, leg=54.1202, rel=1.33e-03 |
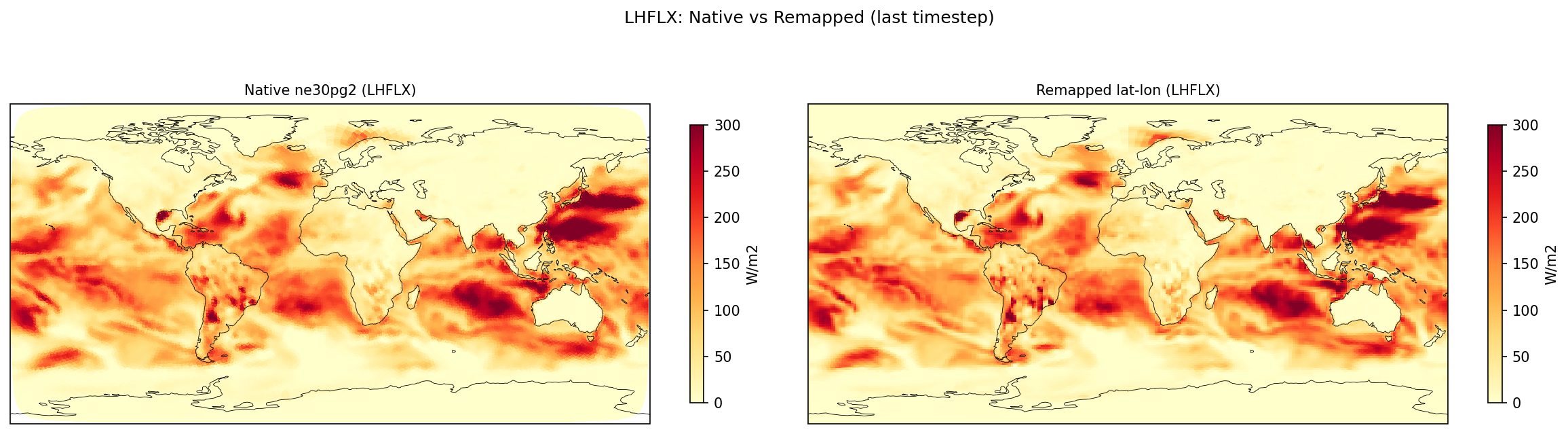
| LHFLX | PASS | fme=80.3688, leg=80.2388, rel=1.62e-03 |
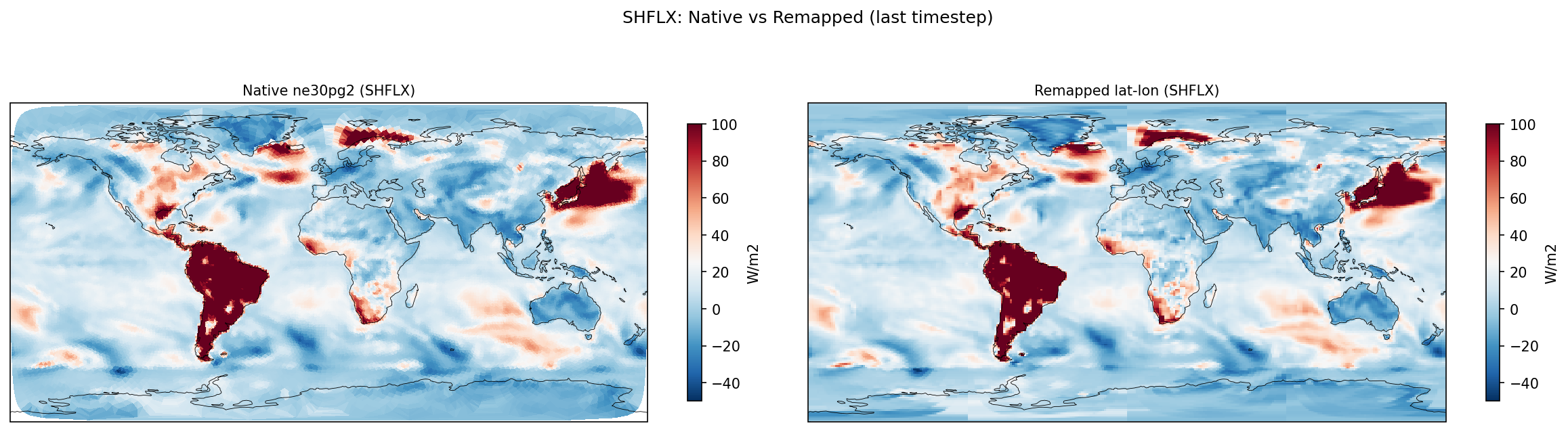
| SHFLX | PASS | fme=17.4038, leg=17.6629, rel=1.47e-02 |
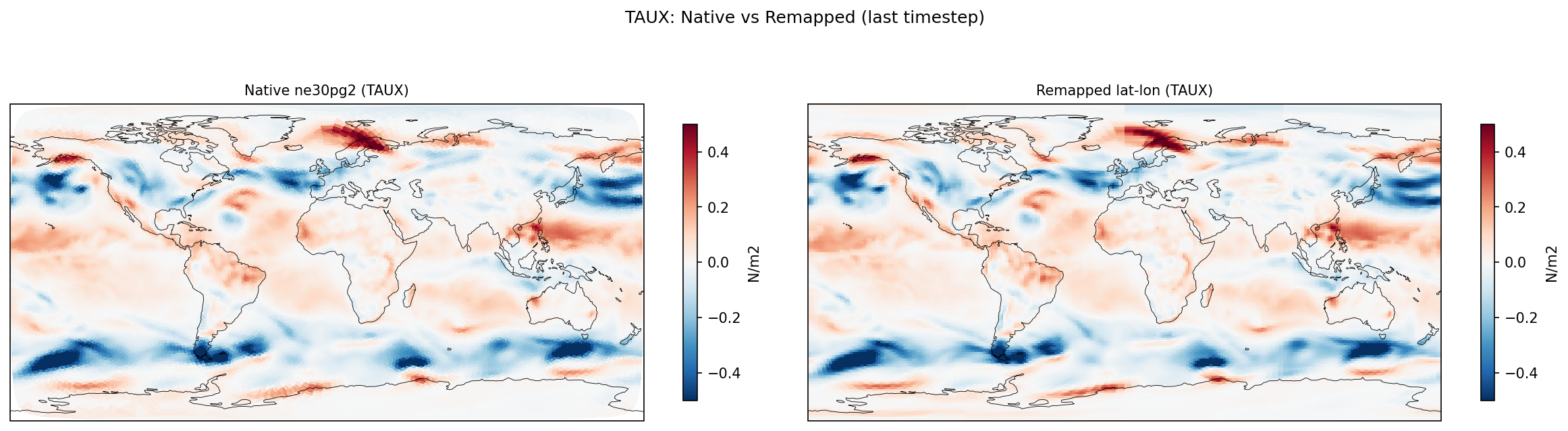
| TAUX | DIFF | fme=-0.004732, leg=-0.00750318, rel=3.69e-01 |
| TAUY | DIFF | fme=0.00832446, leg=0.00850632, rel=2.14e-02 |
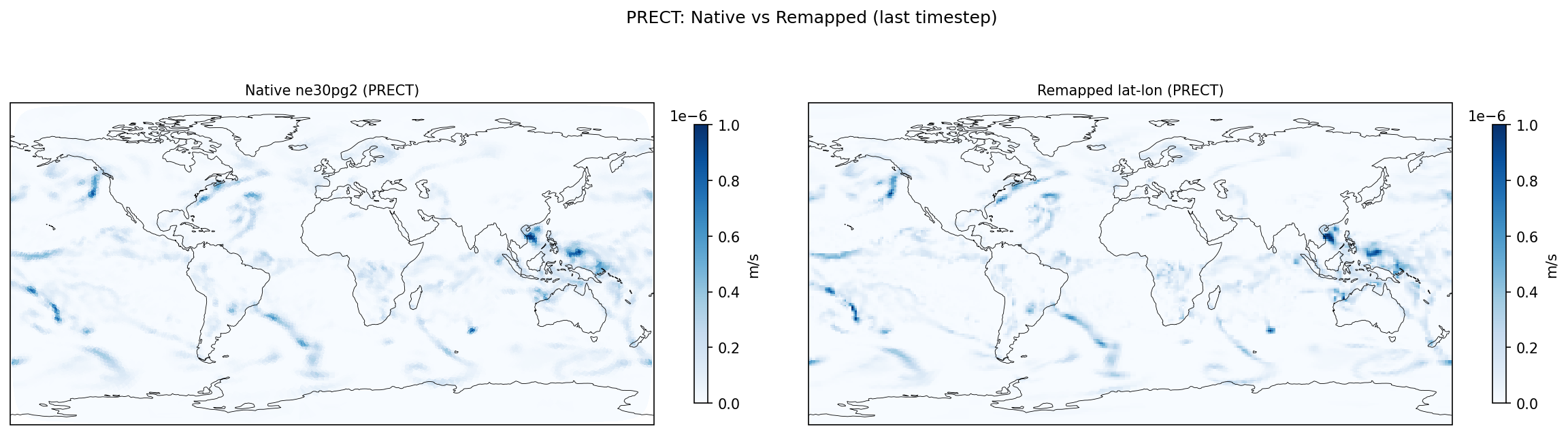
| PRECT | PASS | fme=3.17636e-08, leg=3.21008e-08, rel=1.05e-02 |
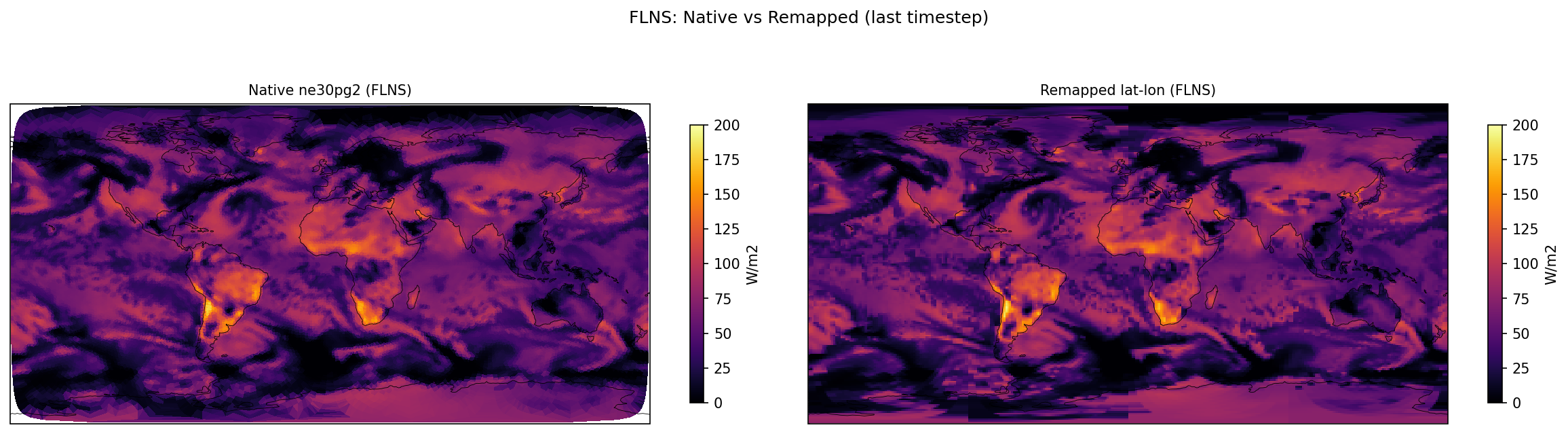
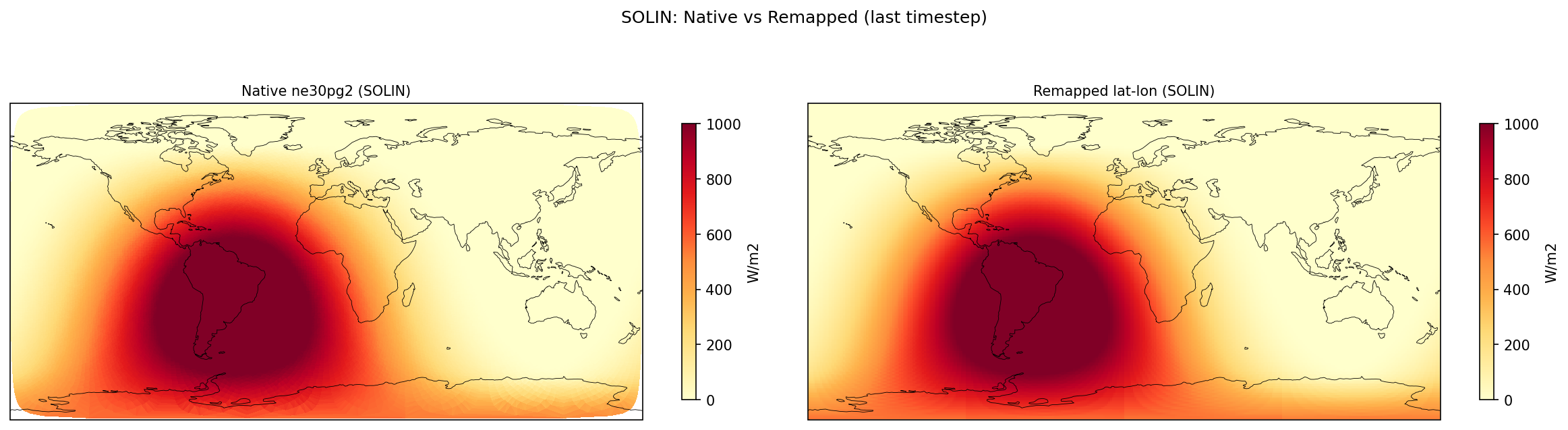
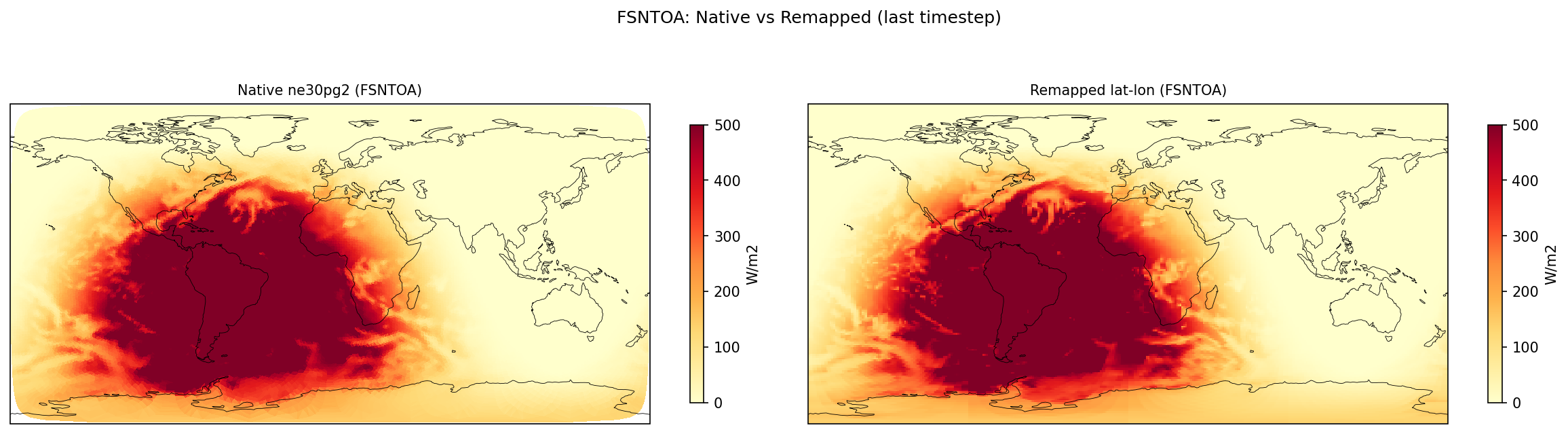
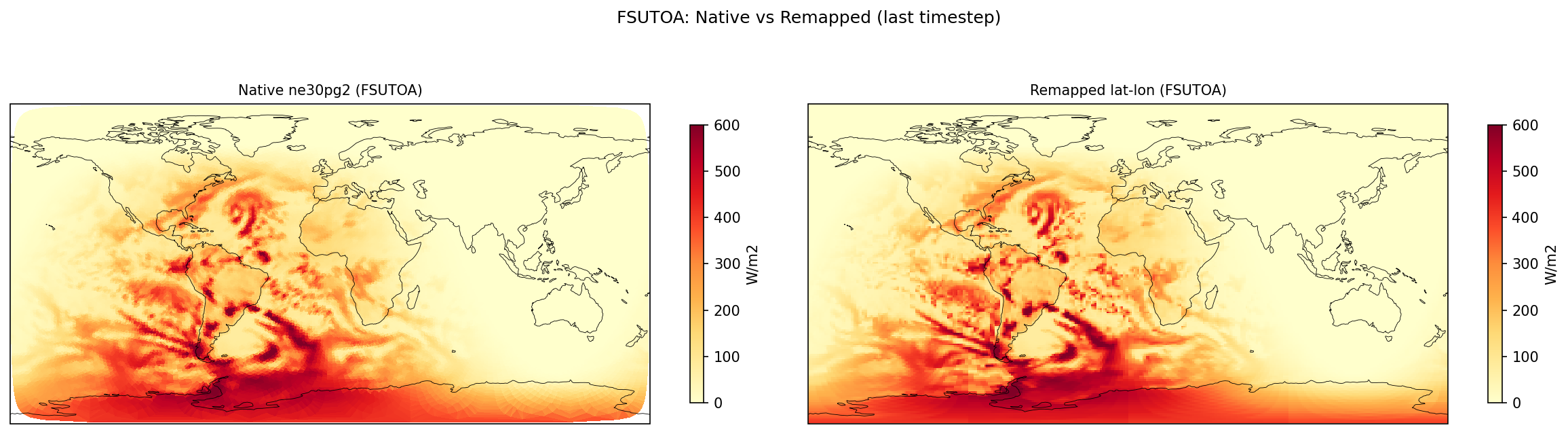
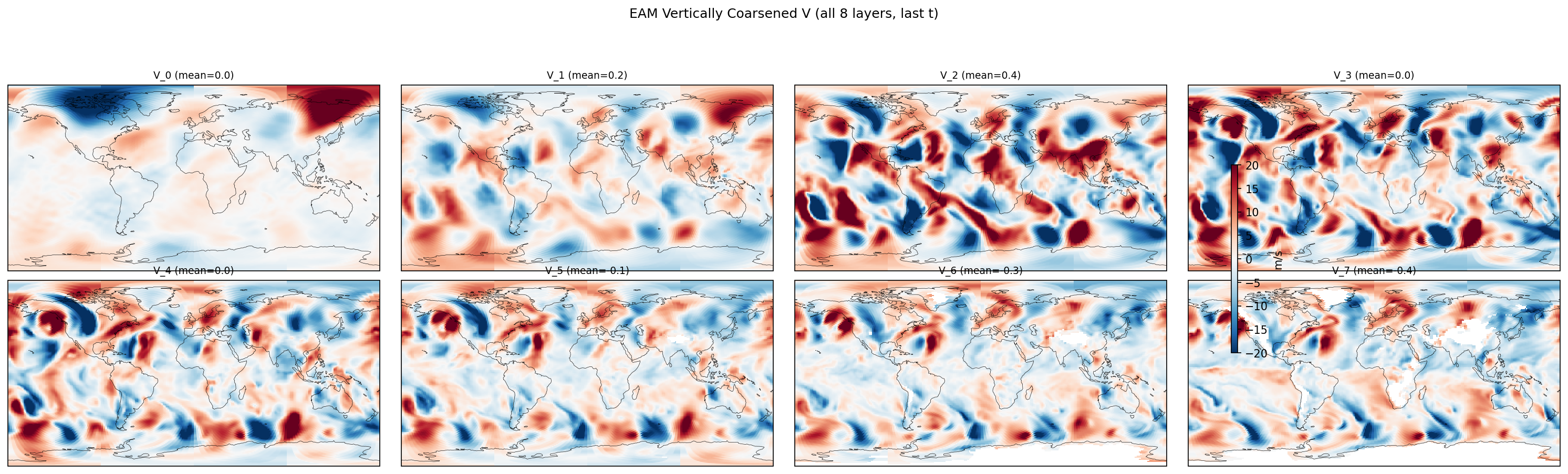
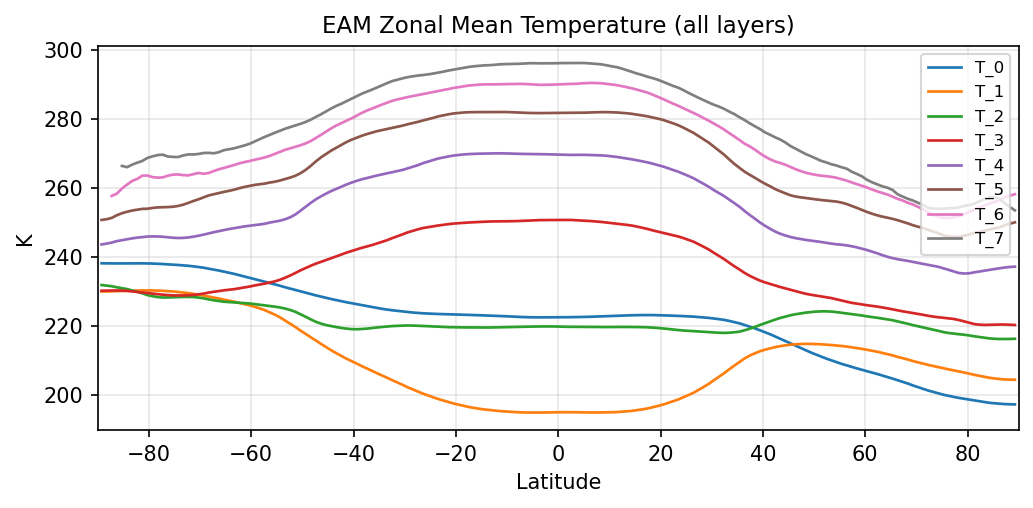
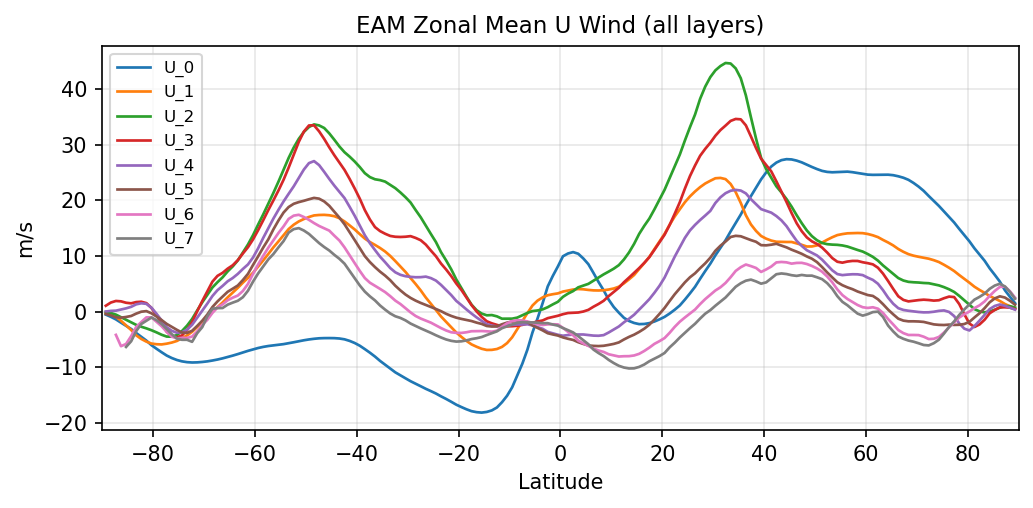
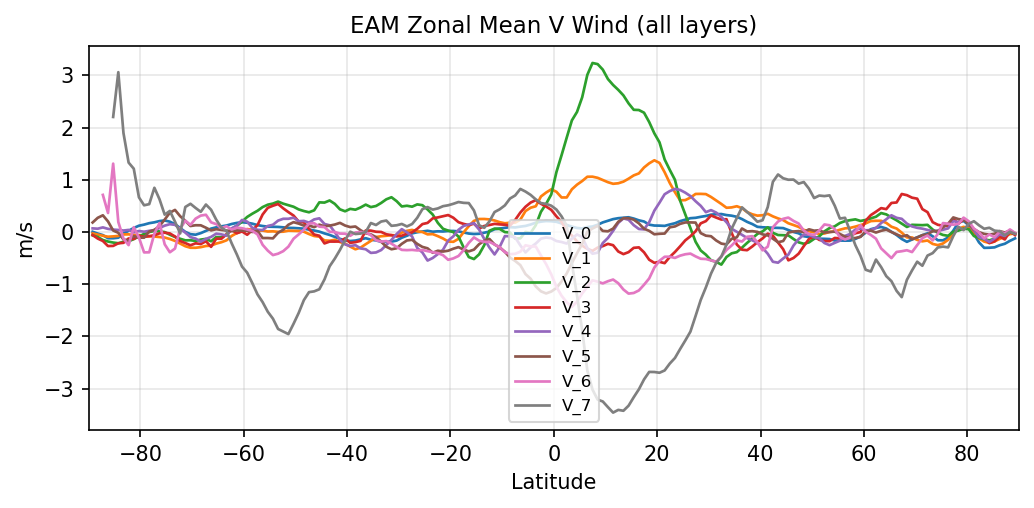
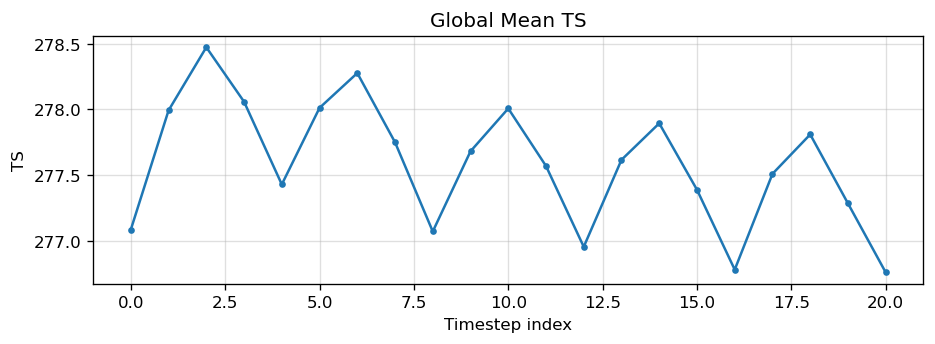
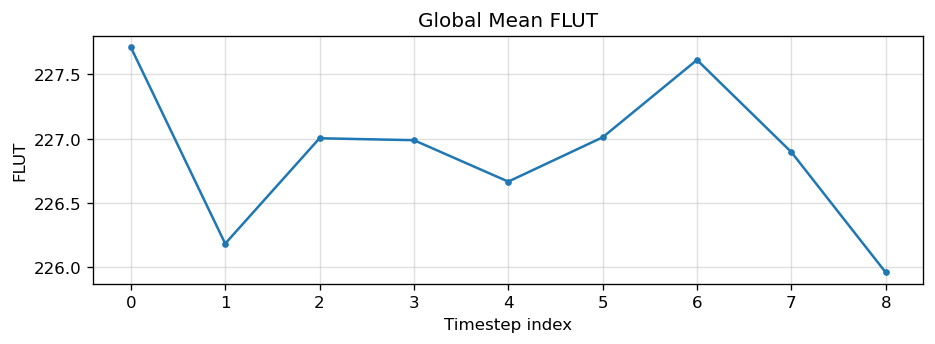
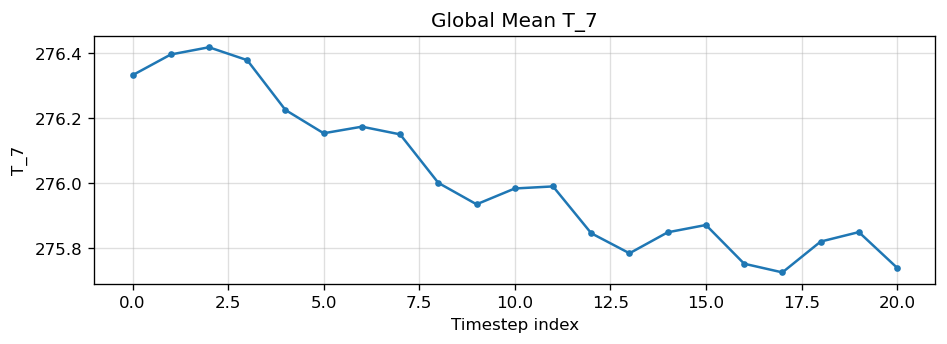
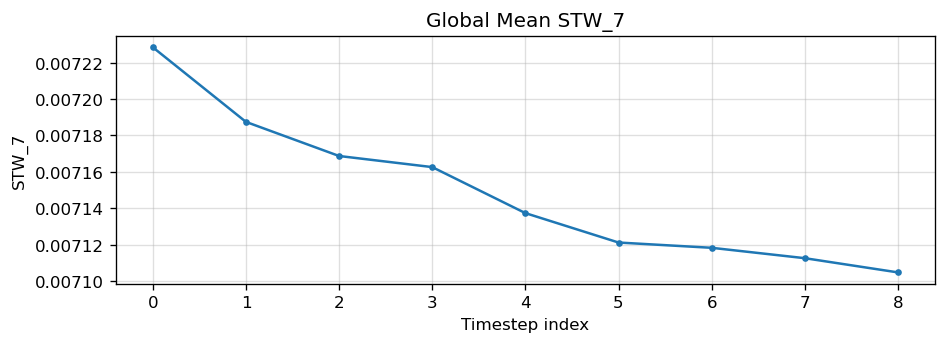
Diagnostic Figures
EAM
Comparisons
Cross-Verify
Performance Timing
| ***** | GLOBAL | STATISTICS | ( | 1024 | MPI | TASKS) | ***** | ||||||
| name | on | processes | threads | count | walltotal | wallmax | (proc | thrd | ) | wallmin | (proc | thrd | ) |
| "CPL:INIT" | - | 1024 | 1024 | 1.024000e+03 | 7.488779e+04 | 76.070 | ( | 610 | 0) | 69.766 | ( | 983 | 0) |
| "CPL:cime_pre_init1" | - | 1024 | 1024 | 1.024000e+03 | 4.429327e+03 | 7.263 | ( | 610 | 0) | 0.959 | ( | 983 | 0) |
| "CPL:mpi_init" | - | 1024 | 1024 | 1.024000e+03 | 4.310037e+03 | 7.147 | ( | 553 | 0) | 0.843 | ( | 983 | 0) |
| "CPL:ESMF_Initialize" | - | 1024 | 1024 | 1.024000e+03 | 2.513084e+00 | 0.004 | ( | 953 | 0) | 0.000 | ( | 688 | 0) |
| "CPL:cime_pre_init2" | - | 1024 | 1024 | 1.024000e+03 | 4.093555e+02 | 0.403 | ( | 374 | 0) | 0.396 | ( | 131 | 0) |
| "CPL:seq_infodata_init" | - | 1024 | 1024 | 1.024000e+03 | 6.452165e+01 | 0.133 | ( | 379 | 0) | 0.002 | ( | 746 | 0) |
| "CPL:seq_timemgr_clockInit" | - | 1024 | 1024 | 1.024000e+03 | 4.781367e+00 | 0.008 | ( | 135 | 0) | 0.001 | ( | 897 | 0) |
| "CPL:cime_init" | - | 1024 | 1024 | 1.024000e+03 | 7.004605e+04 | 68.410 | ( | 0 | 0) | 68.402 | ( | 406 | 0) |
| "CPL:init_comps" | - | 1024 | 1024 | 1.024000e+03 | 5.926836e+04 | 57.887 | ( | 121 | 0) | 57.875 | ( | 168 | 0) |
| "CPL:comp_init_pre_all" | - | 1024 | 1024 | 1.024000e+03 | 2.891074e-01 | 0.002 | ( | 871 | 0) | 0.000 | ( | 939 | 0) |
| "comp_init_cc_iac" | - | 1024 | 1024 | 1.024000e+03 | 2.396527e+00 | 0.006 | ( | 742 | 0) | 0.000 | ( | 501 | 0) |
| "z_i:comp_init" | - | 1024 | 1024 | 1.024000e+03 | 8.589628e-01 | 0.002 | ( | 701 | 0) | 0.000 | ( | 87 | 0) |
| "CPL:comp_init_cc_atm" | - | 1024 | 1024 | 1.024000e+03 | 3.033496e+04 | 29.628 | ( | 1 | 0) | 29.623 | ( | 815 | 0) |
| "a_i:comp_init" | - | 1024 | 1024 | 2.048000e+03 | 3.517122e+04 | 34.483 | ( | 25 | 0) | 34.246 | ( | 137 | 0) |
| "a_i:cam_init" | - | 1024 | 1024 | 1.024000e+03 | 3.007682e+04 | 29.374 | ( | 937 | 0) | 29.369 | ( | 293 | 0) |
| "a_i:cam_initfiles_open" | - | 1024 | 1024 | 1.024000e+03 | 3.328119e+02 | 0.327 | ( | 947 | 0) | 0.322 | ( | 256 | 0) |
| "a_i:cam_initial" | - | 1024 | 1024 | 1.024000e+03 | 6.756930e+03 | 6.600 | ( | 1 | 0) | 6.597 | ( | 916 | 0) |
| "a_i:dyn_init1" | - | 1024 | 1024 | 1.024000e+03 | 6.018509e+02 | 1.653 | ( | 296 | 0) | 0.176 | ( | 542 | 0) |
| "a_i:hycoef_init" | - | 1024 | 1024 | 1.024000e+03 | 6.547680e+01 | 0.074 | ( | 833 | 0) | 0.046 | ( | 32 | 0) |
| "a_i:prim_init1" | - | 1024 | 1024 | 1.024000e+03 | 7.456527e+01 | 0.091 | ( | 56 | 0) | 0.058 | ( | 787 | 0) |
| "a_i:repro_sum_finalsum" | - | 1024 | 1024 | 9.011200e+04 | 4.052332e-02 | 0.000 | ( | 132 | 0) | 0.000 | ( | 655 | 0) |
| "a_i:compose_init" | - | 1024 | 1024 | 1.024000e+03 | 2.088705e+01 | 0.028 | ( | 278 | 0) | 0.015 | ( | 128 | 0) |
| "a_i:sl_init1" | - | 1024 | 1024 | 1.024000e+03 | 5.235690e+00 | 0.011 | ( | 127 | 0) | 0.001 | ( | 215 | 0) |
| "a_i:phys_grid_init" | - | 1024 | 1024 | 1.024000e+03 | 1.114781e+03 | 1.501 | ( | 608 | 0) | 0.024 | ( | 269 | 0) |
| "a_i:initial_conds" | - | 1024 | 1024 | 1.024000e+03 | 4.970694e+03 | 4.871 | ( | 270 | 0) | 4.843 | ( | 813 | 0) |
| "a_i:PIO:initdecomp" | - | 1024 | 1024 | 6.144000e+03 | 4.233158e+01 | 0.059 | ( | 0 | 0) | 0.035 | ( | 937 | 0) |
| "a_i:PIO:PIOc_initdecomp" | - | 1024 | 1024 | 6.144000e+03 | 4.149433e+01 | 0.058 | ( | 0 | 0) | 0.035 | ( | 937 | 0) |
| "a_i:dyn_init2" | - | 1024 | 1024 | 1.024000e+03 | 5.604318e+01 | 0.063 | ( | 585 | 0) | 0.048 | ( | 238 | 0) |
Reproducibility
- Command
/eagle/E3SMinput/mahf708/scratch/crux/mahf708-e3sm/cime_config/testmods_dirs/allactive/fme_output/verify_eam.py --rundir /eagle/E3SMinput/mahf708/scratch/crux/SMS_Ld2.ne30pg2_r05_IcoswISC30E3r5.WCYCL1850.crux_gnu.allactive-fme_output.20260415_162604_ycj7vh/run --legacy-rundir /eagle/E3SMinput/mahf708/scratch/crux/SMS_Ld2.ne30pg2_r05_IcoswISC30E3r5.WCYCL1850.crux_gnu.allactive-fme_legacy_output.20260415_162554_q16mxg/run --outdir /eagle/E3SMinput/mahf708/scratch/crux/share/fme_output_eam -v- Script
/eagle/E3SMinput/mahf708/scratch/crux/mahf708-e3sm/cime_config/testmods_dirs/allactive/fme_output/verify_eam.py— git:1ecfbf15b7 (dirty)- Run directory
/eagle/E3SMinput/mahf708/scratch/crux/SMS_Ld2.ne30pg2_r05_IcoswISC30E3r5.WCYCL1850.crux_gnu.allactive-fme_output.20260415_162604_ycj7vh/run- Legacy run directory
/eagle/E3SMinput/mahf708/scratch/crux/SMS_Ld2.ne30pg2_r05_IcoswISC30E3r5.WCYCL1850.crux_gnu.allactive-fme_legacy_output.20260415_162554_q16mxg/run- Host
mahf708@x1000c6s1b0n0- Generated
- 2026-04-15 19:09:12
user_nl_eam (FME)
! FME (Full Model Emulation) online output processing -- EAM
!
! Production configuration for ACE/Samudra training data generation.
!
! Vertical coarsening uses EAM L80 interface pressures matching the ACE
! offline coarsening indices [0,25,38,46,52,56,61,69,80] (see:
! github.com/ai2cm/ace/blob/exp/e3sm/scripts/data_process/configs/
! e3smv3-coupled-atm-1deg.yaml).
! Derived from hyai/hybi in eami_mam4_Linoz_ne30np4_L80_c20231010.nc.
empty_htapes = .true.
avgflag_pertape = 'I'
nhtfrq = -6
mfilt = 4
! Derived fields (computed at full resolution before vcoarsen)
! STW -> specific_total_water in ACE naming
derived_fld_defs = 'STW=Q+CLDICE+CLDLIQ+RAINQM'
! Vertical coarsening: 8 pressure layers matching ACE L80 coarsening.
! Interface pressures (Pa) from L80 hyai/hybi at indices [0,25,38,46,52,56,61,69,80]:
! 0: 10.0 Pa, 25: 4803.8 Pa, 38: 13913.1 Pa, 46: 26856.3 Pa,
! 52: 43998.3 Pa, 56: 59659.7 Pa, 61: 76854.2 Pa, 69: 90711.8 Pa, 80: 101325.0 Pa
vcoarsen_pbounds = 10.0, 4803.81, 13913.06, 26856.34,
43998.31, 59659.67, 76854.15, 90711.83, 101325.0
vcoarsen_avg_flds = 'T','U','V','STW'
! Tape 1: 6-hourly output.
! State variables are instantaneous; fluxes/tendencies use :A (time-averaged).
fincl1 = 'SOLIN:A', 'PHIS',
'LANDFRAC', 'OCNFRAC', 'ICEFRAC',
'PS', 'TS',
'T_0','T_1','T_2','T_3','T_4','T_5','T_6','T_7',
'STW_0', 'STW_1','STW_2','STW_3','STW_4','STW_5','STW_6','STW_7',
'U_0','U_1','U_2','U_3','U_4','U_5','U_6','U_7',
'V_0','V_1','V_2','V_3','V_4','V_5','V_6','V_7',
'FLUT:A', 'FLNT:A', 'FSNTOA:A', 'FSUTOA:A',
'FLDS:A', 'FLUS:A', 'FSDS:A', 'FSUS:A', 'FSNS:A', 'FLNS:A',
'LHFLX:A','SHFLX:A',
'TAUX:A', 'TAUY:A',
'PRECT:A',
'DTENDTTW'
horiz_remap_file(1) = '/eagle/E3SMinput/mahf708/fme_maps/map_ne30pg2_to_gaussian_180by360_shifted.nc'
user_nl_eam (Legacy)
cosp_lite = .true.
empty_htapes = .true.
avgflag_pertape = 'I'
nhtfrq = -6
mfilt = 4
fincl1 = 'T','Q','U','V','PS','PHIS',
'CLDLIQ','CLDICE','RAINQM',
'TMQ','TGCLDLWP','TGCLDIWP',
'TEFIX','TS','QREFHT','U10','TREFHT',
'PSL','LANDFRAC','OCNFRAC','ICEFRAC',
'FSDS:A','FLDS:A','FSNS:A','FLNS:A','SOLIN:A',
'FSNTOA:A','FSUTOA:A','FLUT:A','FLNT:A',
'FSUS:A','FLUS:A',
'LHFLX:A','QFLX:A',
'SHFLX:A','PRECT:A','PRECC:A','PRECSL:A','PRECSC:A','TAUX:A','TAUY:A'
verify_eam.py (3064 lines)
1 #!/usr/bin/env python3 2 """ 3 Verify EAM FME online output (vcoarsen, derived fields) and cross-verify 4 against legacy output. Produces HTML dashboard with BFB identity tests, 5 offline vcoarsen comparison, and diagnostic figures. 6 7 Usage: 8 python verify_eam.py --rundir /path/to/FME/RUNDIR --outdir /path/to/figs 9 python verify_eam.py --rundir /path/to/FME/RUNDIR --legacy-rundir /path/to/LEGACY/RUNDIR --outdir /path/to/figs 10 """ 11 12 import argparse 13 import os 14 import sys 15 import glob 16 import textwrap 17 import time as _time 18 from datetime import datetime 19 from pathlib import Path 20 21 import numpy as np 22 23 # -- optional imports ---------------------------------------------------------- 24 try: 25 import xarray as xr 26 HAS_XR = True 27 except ImportError: 28 HAS_XR = False 29 30 try: 31 import matplotlib 32 matplotlib.use("Agg") 33 import matplotlib.pyplot as plt 34 import matplotlib.colors as mcolors 35 HAS_MPL = True 36 except ImportError: 37 HAS_MPL = False 38 39 try: 40 import cartopy.crs as ccrs 41 import cartopy.feature as cfeature 42 HAS_CARTOPY = True 43 except ImportError: 44 HAS_CARTOPY = False 45 46 try: 47 from netCDF4 import Dataset as NC4Dataset 48 HAS_NC4 = True 49 except ImportError: 50 HAS_NC4 = False 51 52 # -- ACE atmosphere variable requirements ------------------------------------ 53 # Single source of truth for the ACE-to-EAM variable mapping. 54 # Drives verification, HTML coverage table, and reproducibility tracking. 55 # 56 # The lists below mirror the ACE YAML specification exactly. 57 # EAM field names differ from ACE canonical names in several cases; 58 # ace_to_eam() handles the mapping. 59 60 N_ATM_LAYERS = 8 # ACE uses 8 pressure layers (0-based: T_0..T_7) 61 62 # 3D vertically coarsened fields: ACE base name -> EAM base name 63 ACE_3D_BASE_MAP = { 64 "T": "T", 65 "specific_total_water": "STW", 66 "U": "U", 67 "V": "V", 68 } 69 70 # EAM coarsened base names (used by check_eam and comparison figures) 71 ACE_ATM_3D_COARSENED = list(ACE_3D_BASE_MAP.values()) # ["T", "STW", "U", "V"] 72 73 # 2D / scalar field mapping: ACE canonical name -> EAM FME output name 74 ACE_2D_MAP = { 75 "land_fraction": "LANDFRAC", 76 "ocean_fraction": "OCNFRAC", 77 "sea_ice_fraction": "ICEFRAC", 78 "PHIS": "PHIS", 79 "SOLIN": "SOLIN", 80 "PS": "PS", 81 "TS": "TS", 82 "LHFLX": "LHFLX", 83 "SHFLX": "SHFLX", 84 "surface_precipitation_rate": "PRECT", 85 "surface_upward_longwave_flux": "FLUS", 86 "FLUT": "FLUT", 87 "FLDS": "FLDS", 88 "FSDS": "FSDS", 89 "surface_upward_shortwave_flux": "FSUS", 90 "top_of_atmos_upward_shortwave_flux": "FSUTOA", 91 "tendency_of_total_water_path_due_to_advection": "DTENDTTW", 92 "TAUX": "TAUX", 93 "TAUY": "TAUY", 94 } 95 96 # ACE variable lists (mirror the YAML specification) 97 ACE_FORCING_NAMES = ["SOLIN"] 98 99 ACE_IN_NAMES = ([ 100 "land_fraction", "ocean_fraction", "sea_ice_fraction", 101 "PHIS", "SOLIN", "PS", "TS", 102 ] + [f"T_{k}" for k in range(N_ATM_LAYERS)] 103 + [f"specific_total_water_{k}" for k in range(N_ATM_LAYERS)] 104 + [f"U_{k}" for k in range(N_ATM_LAYERS)] 105 + [f"V_{k}" for k in range(N_ATM_LAYERS)]) 106 107 ACE_OUT_NAMES = ([ 108 "PS", "TS", 109 ] + [f"T_{k}" for k in range(N_ATM_LAYERS)] 110 + [f"specific_total_water_{k}" for k in range(N_ATM_LAYERS)] 111 + [f"U_{k}" for k in range(N_ATM_LAYERS)] 112 + [f"V_{k}" for k in range(N_ATM_LAYERS)] 113 + [ 114 "LHFLX", "SHFLX", 115 "surface_precipitation_rate", 116 "surface_upward_longwave_flux", 117 "FLUT", "FLDS", "FSDS", 118 "surface_upward_shortwave_flux", 119 "top_of_atmos_upward_shortwave_flux", 120 "tendency_of_total_water_path_due_to_advection", 121 "TAUX", "TAUY", 122 ]) 123 124 125 def ace_to_eam(ace_name): 126 """Map an ACE canonical variable name to its EAM FME output name.""" 127 if ace_name in ACE_2D_MAP: 128 return ACE_2D_MAP[ace_name] 129 # 3D indexed variables (e.g. specific_total_water_3 -> STW_3) 130 for ace_base in sorted(ACE_3D_BASE_MAP, key=len, reverse=True): 131 prefix = ace_base + "_" 132 if ace_name.startswith(prefix): 133 suffix = ace_name[len(ace_base):] 134 return ACE_3D_BASE_MAP[ace_base] + suffix 135 return ace_name 136 137 138 # Physical range checks: (vmin, vmax) 139 # Ranges are deliberately generous to avoid false positives during spinup. 140 RANGE_CHECKS = { 141 "T": (150, 350), 142 "Q": (0, 0.05), 143 "PS": (40000, 110000), 144 "TS": (200, 340), 145 "PHIS": (-1000, 60000), 146 } 147 148 # -- Vcoarsen constants ------------------------------------------------------- 149 GRAVIT = 9.80616 # m/s2, matches EAM physconst 150 P0 = 100000.0 # Pa, reference pressure for hybrid coordinates 151 VCOARSEN_PBOUNDS = np.array([ 152 10.0, 4803.81, 13913.06, 26856.34, 153 43998.31, 59659.67, 76854.15, 90711.83, 101325.0 154 ]) # 9 interfaces -> 8 layers, matching ACE L80 indices [0,25,38,46,52,56,61,69,80] 155 N_VCOARSEN_LAYERS = len(VCOARSEN_PBOUNDS) - 1 # 8 156 157 # Fields that appear in BOTH fme_output and fme_legacy_output for BFB identity checks 158 # Instantaneous (no :A suffix) 159 BFB_INST_FIELDS = ["PS", "TS", "PHIS", "LANDFRAC", "OCNFRAC", "ICEFRAC"] 160 # Averaged (:A suffix in both cases) 161 BFB_AVG_FIELDS = ["SOLIN", "FSNTOA", "FSUTOA", "FLUT", "FLNT", 162 "FLDS", "FLUS", "FSDS", "FSUS", "FSNS", "FLNS", 163 "LHFLX", "SHFLX", "TAUX", "TAUY", "PRECT"] 164 165 # FME vcoarsen fields -> legacy raw source (for offline recomputation) 166 # FME name -> (legacy fields to sum for derived, then vcoarsen) 167 FME_TO_LEGACY = { 168 "T": ["T"], 169 "U": ["U"], 170 "V": ["V"], 171 "STW": ["Q", "CLDICE", "CLDLIQ", "RAINQM"], 172 } 173 174 175 # -- Offline vcoarsen / column integration ------------------------------------ 176 177 def compute_pint(ps, hyai, hybi): 178 """Compute interface pressures from hybrid coordinates. 179 180 Parameters 181 ---------- 182 ps : ndarray, shape (ncol,) 183 Surface pressure in Pa. 184 hyai, hybi : ndarray, shape (nlev+1,) 185 Hybrid A and B coefficients (interface levels, top to bottom). 186 187 Returns 188 ------- 189 pint : ndarray, shape (ncol, nlev+1) 190 Interface pressures in Pa. 191 """ 192 # pint(i, k) = hyai(k) * P0 + hybi(k) * ps(i) 193 return hyai[np.newaxis, :] * P0 + hybi[np.newaxis, :] * ps[:, np.newaxis] 194 195 196 def compute_pdel(pint): 197 """Compute layer thickness in Pa from interface pressures. 198 199 Parameters 200 ---------- 201 pint : ndarray, shape (ncol, nlev+1) 202 203 Returns 204 ------- 205 pdel : ndarray, shape (ncol, nlev) 206 """ 207 return pint[:, 1:] - pint[:, :-1] 208 209 210 def offline_vcoarsen_avg(field, pint, pbounds=None): 211 """Overlap-weighted vertical averaging onto coarsened pressure layers. 212 213 Implements the same algorithm as shr_vcoarsen_avg in shr_vcoarsen_mod.F90. 214 For each target layer [pb_top, pb_bot], computes the pressure-weighted mean 215 of all model levels that overlap with it. 216 217 Parameters 218 ---------- 219 field : ndarray, shape (ncol, nlev) 220 Full-resolution field values. 221 pint : ndarray, shape (ncol, nlev+1) 222 Interface pressures from compute_pint. 223 pbounds : ndarray, shape (nlayers+1,), optional 224 Target layer pressure boundaries. Defaults to VCOARSEN_PBOUNDS. 225 226 Returns 227 ------- 228 coarsened : ndarray, shape (ncol, nlayers) 229 Coarsened field values. 230 """ 231 if pbounds is None: 232 pbounds = VCOARSEN_PBOUNDS 233 nlayers = len(pbounds) - 1 234 ncol, nlev = field.shape 235 236 coarsened = np.zeros((ncol, nlayers)) 237 for layer in range(nlayers): 238 pb_top = pbounds[layer] 239 pb_bot = pbounds[layer + 1] 240 241 weight_sum = np.zeros(ncol) 242 for k in range(nlev): 243 # Overlap between model level k and target layer 244 overlap_top = np.maximum(pb_top, pint[:, k]) 245 overlap_bot = np.minimum(pb_bot, pint[:, k + 1]) 246 overlap = np.maximum(0.0, overlap_bot - overlap_top) 247 248 # Skip NaN and fill values — only average valid data 249 fk = field[:, k] 250 ok = np.isfinite(fk) & (overlap > 0) 251 coarsened[ok, layer] += fk[ok] * overlap[ok] 252 weight_sum[ok] += overlap[ok] 253 254 # Normalize by total overlap of valid levels 255 valid = weight_sum > 0 256 coarsened[valid, layer] /= weight_sum[valid] 257 coarsened[~valid, layer] = np.nan 258 259 return coarsened 260 261 262 def offline_column_integrate(field, pdel): 263 """Column integration: sum(field * pdel / g) over all levels. 264 265 Parameters 266 ---------- 267 field : ndarray, shape (ncol, nlev) 268 pdel : ndarray, shape (ncol, nlev) 269 270 Returns 271 ------- 272 integrated : ndarray, shape (ncol,) 273 """ 274 return np.sum(field * pdel / GRAVIT, axis=1) 275 276 277 # ----------------------------------------------------------------------------- 278 # Utility helpers 279 # ----------------------------------------------------------------------------- 280 281 def find_files(rundir, pattern, exclude=None): 282 """Find files matching glob pattern, optionally excluding a substring. 283 284 Parameters 285 ---------- 286 rundir : str 287 Directory to search in. 288 pattern : str 289 Glob pattern relative to rundir. 290 exclude : str or None 291 If set, exclude any file whose basename contains this substring. 292 293 Returns 294 ------- 295 list of str 296 Sorted list of matching file paths. 297 """ 298 hits = sorted(glob.glob(os.path.join(rundir, pattern))) 299 # Always exclude .base restart files 300 hits = [f for f in hits if not f.endswith('.base')] 301 if exclude: 302 hits = [f for f in hits if exclude not in os.path.basename(f)] 303 return hits 304 305 306 def safe_open(path): 307 """Open a NetCDF file with xarray (preferred) or netCDF4.""" 308 if HAS_XR: 309 try: 310 return xr.open_dataset(path, decode_times=False) 311 except Exception: 312 pass 313 if HAS_NC4: 314 return NC4Dataset(path, "r") 315 raise RuntimeError("Neither xarray nor netCDF4 is available.") 316 317 318 def get_var(ds, name): 319 """Return numpy array from xarray Dataset or netCDF4 Dataset.""" 320 if HAS_XR and isinstance(ds, xr.Dataset): 321 if name in ds: 322 return ds[name].values 323 return None 324 if HAS_NC4 and isinstance(ds, NC4Dataset): 325 if name in ds.variables: 326 return np.array(ds.variables[name][:]) 327 return None 328 return None 329 330 331 def get_dims(ds): 332 """Return dict of dimension names -> sizes.""" 333 if HAS_XR and isinstance(ds, xr.Dataset): 334 return dict(ds.sizes) 335 if HAS_NC4 and isinstance(ds, NC4Dataset): 336 return {d: len(ds.dimensions[d]) for d in ds.dimensions} 337 return {} 338 339 340 def get_varnames(ds): 341 """Return list of variable names in the dataset.""" 342 if HAS_XR and isinstance(ds, xr.Dataset): 343 return list(ds.data_vars) 344 if HAS_NC4 and isinstance(ds, NC4Dataset): 345 return list(ds.variables.keys()) 346 return [] 347 348 349 def close_ds(ds): 350 """Close a dataset if applicable.""" 351 if HAS_XR and isinstance(ds, xr.Dataset): 352 ds.close() 353 elif HAS_NC4 and isinstance(ds, NC4Dataset): 354 ds.close() 355 356 357 def valid_data(arr, fill=1e10): 358 """Return non-fill, finite values. 359 360 The threshold is 1e10 because remapped output can contain interpolated 361 fill artifacts from neighboring land/ocean cells that can reach ~1e14 362 scale while _FillValue is 1e34. No geophysical quantity exceeds 1e10. 363 """ 364 if arr is None: 365 return None 366 flat = arr.ravel().astype(float) 367 mask = (np.abs(flat) < fill) & np.isfinite(flat) 368 return flat[mask] if mask.any() else None 369 370 371 def fill_nan_report(arr, name): 372 """Return a dict with fill/NaN fraction stats for a variable array.""" 373 if arr is None: 374 return {"name": name, "total": 0, "valid": 0, "fill_nan": 0, "pct": 100.0} 375 flat = arr.ravel().astype(float) 376 total = flat.size 377 fill_mask = np.abs(flat) >= 1e10 378 nan_mask = ~np.isfinite(flat) 379 bad = fill_mask | nan_mask 380 n_bad = int(bad.sum()) 381 n_valid = total - n_bad 382 pct = 100.0 * n_bad / total if total > 0 else 0.0 383 return {"name": name, "total": total, "valid": n_valid, 384 "fill_nan": n_bad, "pct": pct} 385 386 387 def check_range(arr, name, vmin=None, vmax=None): 388 v = valid_data(arr) 389 issues = [] 390 if v is None or v.size == 0: 391 issues.append(f" {name}: no valid data") 392 return issues 393 if not np.isfinite(v).all(): 394 issues.append(f" {name}: NaN/Inf present") 395 if vmin is not None and v.min() < vmin: 396 issues.append(f" {name}: min={v.min():.4g} below {vmin}") 397 if vmax is not None and v.max() > vmax: 398 issues.append(f" {name}: max={v.max():.4g} above {vmax}") 399 return issues 400 401 402 def summary_stats(arr): 403 v = valid_data(arr) 404 if v is None or v.size == 0: 405 return "no valid data" 406 return f"mean={v.mean():.4g} min={v.min():.4g} max={v.max():.4g} n={v.size}" 407 408 409 def has_time_zero(ds): 410 """Check if the Time/time dimension has size 0.""" 411 dims = get_dims(ds) 412 for tname in ("Time", "time"): 413 if tname in dims and dims[tname] == 0: 414 return True 415 return False 416 417 418 def last_time_index(arr): 419 """Return the index of the last timestep. 420 421 Uses -1 (last available) to avoid initialization artifacts at t=0. 422 """ 423 if arr is None: 424 return -1 425 if arr.ndim >= 2 and arr.shape[0] > 0: 426 return -1 427 return -1 428 429 430 # ----------------------------------------------------------------------------- 431 # Plotting helpers 432 # ----------------------------------------------------------------------------- 433 434 def savefig(fig, outdir, name, subdir=None): 435 """Save figure, optionally in a subdirectory.""" 436 if subdir: 437 d = os.path.join(outdir, subdir) 438 os.makedirs(d, exist_ok=True) 439 path = os.path.join(d, name) 440 else: 441 path = os.path.join(outdir, name) 442 fig.savefig(path, dpi=120, bbox_inches="tight") 443 plt.close(fig) 444 return path 445 446 447 def _fix_lon(lons): 448 """Shift longitudes from [0,360) to [-180,180) for plotting.""" 449 return (lons + 180) % 360 - 180 450 451 452 def _plot_on_ax(ax, data, lons, lats, cmap, vmin, vmax, is_cartopy): 453 """Plot data on an axis -- tripcolor for 1D, pcolormesh for 2D.""" 454 # Mask fill values so they render as transparent (not clipped to vmax) 455 data = np.where((np.abs(data) < 1e10) & np.isfinite(data), 456 data.astype(float), np.nan) 457 transform = ccrs.PlateCarree() if is_cartopy else None 458 if data.ndim == 2: 459 kw = dict(transform=transform) if is_cartopy else {} 460 im = ax.pcolormesh(lons, lats, data, cmap=cmap, 461 vmin=vmin, vmax=vmax, shading="auto", **kw) 462 else: 463 # Unstructured: use tripcolor for filled triangulation. 464 # Subsample if > 50k points to keep triangulation fast. 465 # Mask triangles that cross the antimeridian (lon span > 180). 466 from matplotlib.tri import Triangulation 467 # Flatten in case caller passed 2D per-column coordinate arrays 468 lons_1d = np.asarray(lons).ravel() 469 lats_1d = np.asarray(lats).ravel() 470 data_1d = np.asarray(data).ravel() 471 lons_fix = _fix_lon(lons_1d) 472 kw = dict(transform=ccrs.PlateCarree()) if is_cartopy else {} 473 max_pts = 50000 474 if data_1d.size > max_pts: 475 idx = np.random.default_rng(42).choice(data_1d.size, max_pts, replace=False) 476 lons_sub, lats_sub, data_sub = lons_fix[idx], lats_1d[idx], data_1d[idx] 477 else: 478 lons_sub, lats_sub, data_sub = lons_fix, lats_1d, data_1d 479 try: 480 tri = Triangulation(lons_sub, lats_sub) 481 # Mask triangles spanning the antimeridian 482 tri_lons = lons_sub[tri.triangles] 483 lon_span = tri_lons.max(axis=1) - tri_lons.min(axis=1) 484 tri.set_mask(lon_span > 180) 485 im = ax.tripcolor(tri, data_sub, cmap=cmap, 486 vmin=vmin, vmax=vmax, **kw) 487 except Exception: 488 # Fallback to scatter if tripcolor fails 489 im = ax.scatter(lons_sub, lats_sub, c=data_sub, s=0.5, cmap=cmap, 490 vmin=vmin, vmax=vmax, **kw) 491 return im 492 493 494 def global_map(data, lons, lats, title, cmap="RdBu_r", vmin=None, vmax=None, 495 outdir=None, fname=None, units="", subdir=None): 496 """ 497 Plot a global filled map. data is 2-D (lat x lon) or 1-D (nCells) on an 498 unstructured grid (tripcolor with scatter fallback). 499 """ 500 if not HAS_MPL: 501 return None 502 503 fig = plt.figure(figsize=(10, 5)) 504 is_cartopy = HAS_CARTOPY 505 if is_cartopy: 506 ax = fig.add_subplot(1, 1, 1, projection=ccrs.PlateCarree()) 507 ax.add_feature(cfeature.COASTLINE, linewidth=0.5) 508 ax.set_global() 509 else: 510 ax = fig.add_subplot(1, 1, 1) 511 512 im = _plot_on_ax(ax, data, lons, lats, cmap, vmin, vmax, is_cartopy) 513 plt.colorbar(im, ax=ax, shrink=0.6 if is_cartopy else 0.8, label=units) 514 515 if not is_cartopy: 516 ax.set_xlabel("lon"); ax.set_ylabel("lat") 517 518 ax.set_title(title, fontsize=11) 519 if outdir and fname: 520 return savefig(fig, outdir, fname, subdir=subdir) 521 plt.show() 522 return None 523 524 525 def latlon_map(ds, varname, title, cmap="RdBu_r", vmin=None, vmax=None, 526 outdir=None, fname=None, units="", subdir=None): 527 """Plot a variable from a remapped (lon, lat, Time) dataset. 528 529 Uses the LAST timestep to avoid initialization artifacts. 530 """ 531 if not HAS_MPL: 532 return None 533 534 arr = get_var(ds, varname) 535 if arr is None: 536 return None 537 # Use last timestep if time dimension present 538 if arr.ndim == 3: 539 data = arr[-1, :, :] 540 elif arr.ndim == 2: 541 data = arr 542 else: 543 return None 544 545 lon = get_var(ds, "lon") 546 lat = get_var(ds, "lat") 547 if lon is None or lat is None: 548 return None 549 550 if lon.ndim == 1 and lat.ndim == 1: 551 lons, lats = np.meshgrid(lon, lat) 552 else: 553 lons, lats = lon, lat 554 555 return global_map(data, lons, lats, title, cmap=cmap, vmin=vmin, vmax=vmax, 556 outdir=outdir, fname=fname, units=units, subdir=subdir) 557 558 559 def layer_profiles(data3d, label, outdir, fname, ylabel="Level index", subdir=None): 560 """Plot global-mean vertical profile from (nlev x ncells) array.""" 561 if not HAS_MPL: 562 return 563 means = [] 564 for lev in range(data3d.shape[0]): 565 v = valid_data(data3d[lev]) 566 means.append(v.mean() if v is not None and v.size > 0 else np.nan) 567 fig, ax = plt.subplots(figsize=(4, 5)) 568 ax.plot(means, np.arange(len(means)), "o-") 569 ax.invert_yaxis() 570 ax.set_xlabel(label) 571 ax.set_ylabel(ylabel) 572 ax.set_title(f"Global-mean vertical profile: {label}") 573 ax.grid(True, alpha=0.4) 574 savefig(fig, outdir, fname, subdir=subdir) 575 576 577 def time_series(values, times_label, title, ylabel, outdir, fname, subdir=None): 578 """Plot a simple time series.""" 579 if not HAS_MPL: 580 return 581 fig, ax = plt.subplots(figsize=(8, 3)) 582 ax.plot(values, ".-") 583 ax.set_xlabel(times_label) 584 ax.set_ylabel(ylabel) 585 ax.set_title(title) 586 ax.grid(True, alpha=0.4) 587 plt.tight_layout() 588 savefig(fig, outdir, fname, subdir=subdir) 589 590 591 def multi_panel_maps(panels, suptitle, outdir, fname, ncols=4, 592 cmap="RdBu_r", vmin=None, vmax=None, units=""): 593 """Plot a grid of small-multiple global maps. 594 595 Parameters 596 ---------- 597 panels : list of (data, lons, lats, subtitle) tuples 598 data can be 2D (lat x lon) for pcolormesh or 1D (nCells) for tripcolor. 599 """ 600 if not HAS_MPL or not panels: 601 return None 602 603 n = len(panels) 604 nrows = (n + ncols - 1) // ncols 605 fig_w = 5 * ncols 606 fig_h = 2.8 * nrows + 0.6 607 is_cartopy = HAS_CARTOPY 608 609 fig = plt.figure(figsize=(fig_w, fig_h)) 610 axes = [] 611 for idx in range(n): 612 kw = dict(projection=ccrs.PlateCarree()) if is_cartopy else {} 613 ax = fig.add_subplot(nrows, ncols, idx + 1, **kw) 614 axes.append(ax) 615 616 im = None 617 for idx, (data, lons, lats, subtitle) in enumerate(panels): 618 ax = axes[idx] 619 if is_cartopy: 620 ax.add_feature(cfeature.COASTLINE, linewidth=0.3) 621 ax.set_global() 622 im = _plot_on_ax(ax, data, lons, lats, cmap, vmin, vmax, is_cartopy) 623 ax.set_title(subtitle, fontsize=9) 624 625 if im is not None: 626 fig.colorbar(im, ax=axes, shrink=0.5, label=units, pad=0.03, aspect=30) 627 fig.suptitle(suptitle, fontsize=12, y=1.01) 628 import warnings 629 with warnings.catch_warnings(): 630 warnings.simplefilter("ignore") 631 plt.tight_layout(rect=[0, 0, 1, 0.97]) 632 path = os.path.join(outdir, fname) 633 fig.savefig(path, dpi=150, bbox_inches="tight") 634 plt.close(fig) 635 return path 636 637 638 def zonal_mean_plot(profiles, lat, title, outdir, fname, ylabel=""): 639 """Plot zonal mean profiles (latitude vs value) for multiple variables. 640 641 Parameters 642 ---------- 643 profiles : list of (zmean_1d, label) tuples 644 zmean_1d is a 1D array of length nlat (zonal mean values). 645 lat : 1D array 646 Latitude values. 647 """ 648 if not HAS_MPL or not profiles: 649 return None 650 651 fig, ax = plt.subplots(figsize=(7, 3.5)) 652 has_data = False 653 for zmean, label in profiles: 654 if zmean is None: 655 continue 656 ax.plot(lat, zmean, label=label, linewidth=1.3) 657 has_data = True 658 659 if not has_data: 660 plt.close(fig) 661 return None 662 663 ax.set_xlabel("Latitude") 664 ax.set_ylabel(ylabel) 665 ax.set_title(title, fontsize=11) 666 ax.legend(fontsize=8, loc="best") 667 ax.grid(True, alpha=0.3) 668 ax.set_xlim(-90, 90) 669 plt.tight_layout() 670 path = os.path.join(outdir, fname) 671 fig.savefig(path, dpi=150, bbox_inches="tight") 672 plt.close(fig) 673 return path 674 675 676 def _compute_zonal_mean(arr, fill_thresh=1e10): 677 """Compute zonal mean from (lat, lon) array, masking fill values.""" 678 if arr is None: 679 return None 680 if arr.ndim == 3: 681 data = arr[-1, :, :] 682 elif arr.ndim == 2: 683 data = arr 684 else: 685 return None 686 masked = np.where(np.abs(data) < fill_thresh, data, np.nan) 687 return np.nanmean(masked, axis=1) 688 689 690 def fill_mask_map(data, lons, lats, title, outdir, fname): 691 """Plot a binary map showing valid (blue) vs fill/NaN (red) cells.""" 692 if not HAS_MPL: 693 return None 694 mask = ((np.abs(data) >= 1e10) | ~np.isfinite(data)).astype(float) 695 cmap = mcolors.ListedColormap(["#3b82f6", "#ef4444"]) 696 bounds = [-0.5, 0.5, 1.5] 697 norm = mcolors.BoundaryNorm(bounds, cmap.N) 698 fig = plt.figure(figsize=(10, 5)) 699 is_cartopy = HAS_CARTOPY 700 if is_cartopy: 701 ax = fig.add_subplot(1, 1, 1, projection=ccrs.PlateCarree()) 702 ax.add_feature(cfeature.COASTLINE, linewidth=0.5) 703 ax.set_global() 704 else: 705 ax = fig.add_subplot(1, 1, 1) 706 transform = ccrs.PlateCarree() if is_cartopy else None 707 if data.ndim == 2: 708 kw = dict(transform=transform) if is_cartopy else {} 709 im = ax.pcolormesh(lons, lats, mask, cmap=cmap, norm=norm, shading="auto", **kw) 710 else: 711 kw = dict(transform=ccrs.PlateCarree()) if is_cartopy else {} 712 lons_fix = _fix_lon(lons) 713 im = ax.scatter(lons_fix, lats, c=mask, s=0.5, cmap=cmap, norm=norm, **kw) 714 cbar = plt.colorbar(im, ax=ax, shrink=0.6 if is_cartopy else 0.8, ticks=[0, 1]) 715 cbar.ax.set_yticklabels(["Valid", "Fill/NaN"]) 716 ax.set_title(title, fontsize=11) 717 return savefig(fig, outdir, fname) 718 719 720 def side_by_side_comparison(data1, lons1, lats1, title1, 721 data2, lons2, lats2, title2, 722 suptitle, outdir, fname, 723 cmap="RdBu_r", vmin=None, vmax=None, units=""): 724 """Two global maps side by side -- native tripcolor on left, remapped pcolormesh on right.""" 725 if not HAS_MPL: 726 return None 727 728 is_cartopy = HAS_CARTOPY 729 fig = plt.figure(figsize=(16, 4.5)) 730 im = None 731 for idx, (data, lons, lats, title) in enumerate([ 732 (data1, lons1, lats1, title1), 733 (data2, lons2, lats2, title2), 734 ]): 735 kw = dict(projection=ccrs.PlateCarree()) if is_cartopy else {} 736 ax = fig.add_subplot(1, 2, idx + 1, **kw) 737 if is_cartopy: 738 ax.add_feature(cfeature.COASTLINE, linewidth=0.4) 739 ax.set_global() 740 im = _plot_on_ax(ax, data, lons, lats, cmap, vmin, vmax, is_cartopy) 741 plt.colorbar(im, ax=ax, shrink=0.7, label=units) 742 ax.set_title(title, fontsize=10) 743 744 fig.suptitle(suptitle, fontsize=12, y=1.02) 745 plt.tight_layout() 746 path = os.path.join(outdir, fname) 747 fig.savefig(path, dpi=150, bbox_inches="tight") 748 plt.close(fig) 749 return path 750 751 752 # ----------------------------------------------------------------------------- 753 # EAM verification + visualization 754 # ----------------------------------------------------------------------------- 755 756 def check_eam(rundir, outdir, verbose): 757 print("\n=== EAM ===") 758 issues = [] 759 plots = [] 760 fill_reports = [] 761 762 # -- tape 1 (h0): all EAM fields (single-tape layout) ------------------- 763 t1_files = find_files(rundir, "*.eam.h0.*.nc") 764 if not t1_files: 765 print(" SKIP: no *.eam.h0.*.nc files found") 766 else: 767 print(f" h0: found {len(t1_files)} file(s)") 768 ds = safe_open(t1_files[0]) 769 if has_time_zero(ds): 770 print(" h0: time dimension has size 0 -- skipping") 771 close_ds(ds) 772 else: 773 for var in ["PHIS", "PS", "TS", "LHFLX", "SHFLX", "FLDS", "FSDS", 774 "FSNTOA", "FSUTOA", "FLUT", "SOLIN", "FSUS", "FLUS", 775 "ICEFRAC", "LANDFRAC", "OCNFRAC", "TAUX", "TAUY", 776 "PRECT", "DTENDTTW"]: 777 arr = get_var(ds, var) 778 if arr is None: 779 issues.append(f" h0 missing: {var}") 780 else: 781 vmin, vmax = RANGE_CHECKS.get(var, (None, None)) 782 issues += check_range(arr, f"h0/{var}", vmin, vmax) 783 fill_reports.append(fill_nan_report(arr, f"h0/{var}")) 784 if verbose: 785 print(f" tape1/{var}: {summary_stats(arr)}") 786 787 # Maps of key 2D fields on lat-lon grid -- use LAST timestep 788 if HAS_MPL and HAS_XR and isinstance(ds, xr.Dataset): 789 lon_name = next((d for d in ["lon", "ncol", "longitude"] if d in ds.dims), None) 790 lat_name = next((d for d in ["lat", "latitude"] if d in ds.dims), None) 791 for var, cmap, vm, vx, units in [ 792 ("TS", "RdBu_r", 220, 320, "K"), 793 ("LHFLX", "viridis", 0, 300, "W/m2"), 794 ("PRECT", "Blues", 0, 1e-3, "kg/m2/s"), 795 ("ICEFRAC", "Blues", 0, 1, "fraction"), 796 ("FLUT", "inferno", 100, 320, "W/m2"), 797 ("FSDS", "YlOrRd", 0, 500, "W/m2"), 798 ("FSUTOA", "YlOrRd", 0, 500, "W/m2"), 799 ("FLUS", "inferno", 200, 500, "W/m2"), 800 ("FSUS", "YlOrRd", 0, 300, "W/m2"), 801 ("DTENDTTW", "RdBu_r", -5e-4, 5e-4, "kg/m2/s"), 802 ]: 803 arr = get_var(ds, var) 804 if arr is None or arr.size == 0: 805 continue 806 # Last timestep 807 data = arr[-1] if arr.ndim >= 2 and arr.shape[0] > 0 else arr 808 if lat_name and lon_name and data.ndim == 2: 809 lons = ds[lon_name].values 810 lats = ds[lat_name].values 811 if lons.ndim == 1: 812 lons, lats = np.meshgrid(lons, lats) 813 p = global_map(data, lons, lats, f"EAM {var} (h0, last t)", 814 cmap=cmap, vmin=vm, vmax=vx, 815 outdir=outdir, fname=f"eam_h0_{var}.png", 816 units=units) 817 if p: plots.append(p) 818 819 # Fill value mask maps for key fields 820 for fvar in ["T_7", "ICEFRAC"]: 821 farr = get_var(ds, fvar) 822 if farr is None or farr.size == 0: 823 continue 824 fdata = farr[-1] if farr.ndim >= 2 and farr.shape[0] > 0 else farr 825 if lat_name and lon_name and fdata.ndim == 2: 826 p = fill_mask_map(fdata, lons, lats, 827 f"Fill Mask: {fvar} (h0, last t)", 828 outdir, f"eam_h0_{fvar}_fill_mask.png") 829 if p: 830 plots.append(p) 831 832 close_ds(ds) 833 834 # -- vcoarsen layers (on h0 in single-tape config) ----------------------- 835 if t1_files: 836 ds3 = safe_open(t1_files[0]) 837 if not has_time_zero(ds3): 838 for base in ACE_ATM_3D_COARSENED: 839 found_layers = [] 840 for k in range(N_ATM_LAYERS): # 0-based: T_0..T_7 841 vname = f"{base}_{k}" 842 arr = get_var(ds3, vname) 843 if arr is not None: 844 found_layers.append(k) 845 vmin, vmax = RANGE_CHECKS.get(base, (None, None)) 846 issues += check_range(arr, f"h0/{vname}", vmin, vmax) 847 fill_reports.append(fill_nan_report(arr, f"h0/{vname}")) 848 if len(found_layers) == N_ATM_LAYERS: 849 print(f" h0/{base}: all {N_ATM_LAYERS} layers present (0-based)") 850 elif found_layers: 851 print(f" h0/{base}: {len(found_layers)}/{N_ATM_LAYERS} layers found " 852 f"(indices {found_layers})") 853 issues.append(f" h0/{base}: only {len(found_layers)}/{N_ATM_LAYERS} layers") 854 else: 855 alts = [v for v in get_varnames(ds3) 856 if v.startswith(base + "_") or v.startswith(base.upper() + "_")] 857 if alts: 858 print(f" h0/{base}: found alternate names: {alts[:5]}") 859 else: 860 issues.append(f" h0 missing coarsened field: {base}_0..{base}_{N_ATM_LAYERS-1}") 861 862 # Plot T_0 (near-top) and T_7 (near-surface) at last timestep 863 if HAS_MPL and HAS_XR and isinstance(ds3, xr.Dataset): 864 lon_name = next((d for d in ["lon", "ncol", "longitude"] if d in ds3.dims), None) 865 lat_name = next((d for d in ["lat", "latitude"] if d in ds3.dims), None) 866 for vname, label_tag in [("T_0", "near-top"), ("T_7", "near-surface")]: 867 arr = get_var(ds3, vname) 868 if arr is None or arr.size == 0: 869 continue 870 data = arr[-1] if arr.ndim >= 2 and arr.shape[0] > 0 else arr 871 if lat_name and lon_name and data.ndim == 2: 872 lons = ds3[lon_name].values 873 lats = ds3[lat_name].values 874 if lons.ndim == 1: 875 lons, lats = np.meshgrid(lons, lats) 876 p = global_map(data, lons, lats, 877 f"EAM {vname} ({label_tag}, h0, last t)", 878 cmap="RdBu_r", vmin=150, vmax=320, 879 outdir=outdir, fname=f"eam_h0_{vname}.png", 880 units="K") 881 if p: plots.append(p) 882 close_ds(ds3) 883 884 _report("EAM", issues) 885 return issues, plots, fill_reports 886 887 888 def _report(name, issues): 889 if issues: 890 print(f" FAIL ({len(issues)} issues):") 891 for msg in issues: 892 print(msg) 893 else: 894 print(f" PASS") 895 896 897 # ----------------------------------------------------------------------------- 898 # Timing summary 899 # ----------------------------------------------------------------------------- 900 901 def read_timing_summary(rundir): 902 """Read model_timing_stats if present and return summary lines.""" 903 timing_path = os.path.join(rundir, "timing", "model_timing_stats") 904 if not os.path.exists(timing_path): 905 return None 906 907 try: 908 with open(timing_path, "r") as fh: 909 lines = fh.readlines() 910 except Exception: 911 return None 912 913 if not lines: 914 return None 915 916 # Extract key timing info: look for overall model time and component times 917 summary = [] 918 for line in lines: 919 stripped = line.strip() 920 if not stripped or stripped.startswith("#"): 921 continue 922 # Keep header lines and lines with timing data 923 # Typical format: component-name seconds seconds seconds 924 if any(kw in stripped.lower() for kw in [ 925 "total", "atm", "lnd", "ocn", "ice", "cpl", "rof", "glc", "wav", 926 "init", "run", "final", "overall", "model", 927 ]): 928 summary.append(stripped) 929 elif len(summary) == 0: 930 # Keep initial header/description lines 931 summary.append(stripped) 932 933 return summary[:40] if summary else None 934 935 936 # ----------------------------------------------------------------------------- 937 # Radiation budget 938 # ----------------------------------------------------------------------------- 939 940 def compute_radiation_budget(ds, outdir, comp_plots): 941 """Compute and plot TOA/surface radiation budget diagnostics.""" 942 if not HAS_MPL: 943 return 944 945 budget_vars = ["SOLIN", "FSUTOA", "FLUT", "FSDS", "FSUS", "FLDS", "FLUS", 946 "LHFLX", "SHFLX"] 947 gmeans = {} 948 for vn in budget_vars: 949 arr = get_var(ds, vn) 950 if arr is None: 951 return # need all variables 952 data = arr[-1] if arr.ndim >= 2 and arr.shape[0] > 0 else arr 953 v = valid_data(data) 954 if v is None or v.size == 0: 955 return 956 gmeans[vn] = float(v.mean()) 957 958 toa_net = gmeans["SOLIN"] - gmeans["FSUTOA"] - gmeans["FLUT"] 959 sfc_net = (gmeans["FSDS"] - gmeans["FSUS"] + gmeans["FLDS"] - gmeans["FLUS"] 960 - gmeans["LHFLX"] - gmeans["SHFLX"]) 961 962 # Bar chart of components and net imbalances 963 labels = list(budget_vars) + ["TOA net", "SFC net"] 964 values = [gmeans[v] for v in budget_vars] + [toa_net, sfc_net] 965 colors = ["#f59e0b" if v > 0 else "#3b82f6" for v in values] 966 colors[-2] = "#16a34a" # TOA net 967 colors[-1] = "#16a34a" # SFC net 968 969 fig, ax = plt.subplots(figsize=(10, 5)) 970 x = np.arange(len(labels)) 971 ax.bar(x, values, color=colors, edgecolor="white", linewidth=0.5) 972 ax.set_xticks(x) 973 ax.set_xticklabels(labels, rotation=45, ha="right", fontsize=9) 974 ax.set_ylabel("W/m2") 975 ax.set_title("Radiation Budget: Global-Mean Components") 976 ax.axhline(0, color="gray", linewidth=0.5) 977 ax.grid(axis="y", alpha=0.3) 978 for i, val in enumerate(values): 979 ax.text(i, val + (5 if val >= 0 else -12), f"{val:.1f}", 980 ha="center", fontsize=7, color="black") 981 plt.tight_layout() 982 path = os.path.join(outdir, "eam_radiation_budget.png") 983 fig.savefig(path, dpi=150, bbox_inches="tight") 984 plt.close(fig) 985 comp_plots.append(path) 986 print(f" Wrote eam_radiation_budget.png") 987 988 # Global map of TOA net radiation (pixel-by-pixel, last timestep) 989 lon_name = next((d for d in ["lon", "ncol", "longitude"] if d in ds.dims), None) 990 lat_name = next((d for d in ["lat", "latitude"] if d in ds.dims), None) 991 if lon_name and lat_name: 992 solin = get_var(ds, "SOLIN") 993 fsutoa = get_var(ds, "FSUTOA") 994 flut = get_var(ds, "FLUT") 995 s_last = solin[-1] if solin.ndim >= 2 else solin 996 fu_last = fsutoa[-1] if fsutoa.ndim >= 2 else fsutoa 997 fl_last = flut[-1] if flut.ndim >= 2 else flut 998 toa_map = s_last - fu_last - fl_last 999 lons = ds[lon_name].values 1000 lats = ds[lat_name].values 1001 if lons.ndim == 1 and lat_name == "lat": 1002 lons, lats = np.meshgrid(lons, lats) 1003 p = global_map(toa_map, lons, lats, 1004 "TOA Net Radiation (SOLIN - FSUTOA - FLUT, last t)", 1005 cmap="RdBu_r", vmin=-200, vmax=200, 1006 outdir=outdir, fname="eam_toa_net_radiation.png", 1007 units="W/m2") 1008 if p: 1009 comp_plots.append(p) 1010 print(f" Wrote eam_toa_net_radiation.png") 1011 1012 1013 # ----------------------------------------------------------------------------- 1014 # EAM comparison figures (multi-panel and zonal mean) 1015 # ----------------------------------------------------------------------------- 1016 1017 def generate_comparison_figures(rundir, fig_root, all_plots_by_comp): 1018 """Generate multi-panel layer views and zonal mean profiles for EAM coarsened fields.""" 1019 if not HAS_MPL: 1020 return 1021 1022 comp_outdir = os.path.join(fig_root, "comparisons") 1023 os.makedirs(comp_outdir, exist_ok=True) 1024 comp_plots = [] 1025 1026 # --- EAM h0: multi-panel T_0..T_7 and STW_0..STW_7 --- 1027 h0_files = find_files(rundir, "*.eam.h0.*.nc") 1028 if h0_files: 1029 ds3 = safe_open(h0_files[0]) 1030 if not has_time_zero(ds3) and HAS_XR and isinstance(ds3, xr.Dataset): 1031 lon_name = next((d for d in ["lon", "ncol", "longitude"] if d in ds3.dims), None) 1032 lat_name = next((d for d in ["lat", "latitude"] if d in ds3.dims), None) 1033 if lon_name and lat_name: 1034 lons = ds3[lon_name].values 1035 lats = ds3[lat_name].values if lat_name != lon_name else None 1036 if lats is not None: 1037 if lons.ndim == 1 and lat_name == "lat": 1038 lons, lats = np.meshgrid(lons, lats) 1039 1040 for base, cmap, vm, vx, units_str in [ 1041 ("T", "RdYlBu_r", 180, 310, "K"), 1042 ("STW", "YlGnBu", 0, 0.02, "kg/kg"), 1043 ("U", "RdBu_r", -40, 40, "m/s"), 1044 ("V", "RdBu_r", -20, 20, "m/s"), 1045 ]: 1046 panels = [] 1047 for k in range(N_ATM_LAYERS): # 0-based 1048 vname = f"{base}_{k}" 1049 arr = get_var(ds3, vname) 1050 if arr is None or arr.size == 0: 1051 continue 1052 data = arr[-1] if arr.ndim >= 2 and arr.shape[0] > 0 else arr 1053 v = valid_data(data) 1054 stats = f"mean={v.mean():.1f}" if v is not None and v.size > 0 else "" 1055 panels.append((data, lons, lats, 1056 f"{vname} ({stats})")) 1057 1058 if panels: 1059 p = multi_panel_maps( 1060 panels, 1061 f"EAM Vertically Coarsened {base} " 1062 f"(all {N_ATM_LAYERS} layers, last t)", 1063 comp_outdir, 1064 f"eam_h0_{base}_all_layers.png", 1065 ncols=4, cmap=cmap, vmin=vm, vmax=vx, 1066 units=units_str) 1067 if p: 1068 comp_plots.append(p) 1069 print(f" Wrote {os.path.basename(p)}") 1070 1071 # Zonal mean of T layers (only for lat-lon data) 1072 if lat_name == "lat": 1073 lat_1d = ds3["lat"].values 1074 profiles = [] 1075 for k in range(N_ATM_LAYERS): # 0-based 1076 zm = _compute_zonal_mean(get_var(ds3, f"T_{k}")) 1077 if zm is not None: 1078 profiles.append((zm, f"T_{k}")) 1079 if profiles: 1080 p = zonal_mean_plot( 1081 profiles, lat_1d, 1082 "EAM Zonal Mean Temperature (all layers)", 1083 comp_outdir, "eam_h0_T_zonal_mean.png", 1084 ylabel="K") 1085 if p: 1086 comp_plots.append(p) 1087 print(f" Wrote {os.path.basename(p)}") 1088 1089 # Zonal mean of U layers 1090 u_profiles = [] 1091 for k in range(N_ATM_LAYERS): 1092 zm = _compute_zonal_mean(get_var(ds3, f"U_{k}")) 1093 if zm is not None: 1094 u_profiles.append((zm, f"U_{k}")) 1095 if u_profiles: 1096 p = zonal_mean_plot( 1097 u_profiles, lat_1d, 1098 "EAM Zonal Mean U Wind (all layers)", 1099 comp_outdir, "eam_h0_U_zonal_mean.png", 1100 ylabel="m/s") 1101 if p: 1102 comp_plots.append(p) 1103 print(f" Wrote {os.path.basename(p)}") 1104 1105 # Zonal mean of V layers 1106 v_profiles = [] 1107 for k in range(N_ATM_LAYERS): 1108 zm = _compute_zonal_mean(get_var(ds3, f"V_{k}")) 1109 if zm is not None: 1110 v_profiles.append((zm, f"V_{k}")) 1111 if v_profiles: 1112 p = zonal_mean_plot( 1113 v_profiles, lat_1d, 1114 "EAM Zonal Mean V Wind (all layers)", 1115 comp_outdir, "eam_h0_V_zonal_mean.png", 1116 ylabel="m/s") 1117 if p: 1118 comp_plots.append(p) 1119 print(f" Wrote {os.path.basename(p)}") 1120 1121 close_ds(ds3) 1122 1123 # --- Topographic fill validation: PS vs vcoarsen fill mask --- 1124 if h0_files: 1125 ds_ps = safe_open(h0_files[0]) 1126 if not has_time_zero(ds_ps) and HAS_XR and isinstance(ds_ps, xr.Dataset): 1127 ps_arr = get_var(ds_ps, "PS") 1128 lon_name = next((d for d in ["lon", "ncol", "longitude"] if d in ds_ps.dims), None) 1129 lat_name = next((d for d in ["lat", "latitude"] if d in ds_ps.dims), None) 1130 if ps_arr is not None and lon_name and lat_name: 1131 lons = ds_ps[lon_name].values 1132 lats = ds_ps[lat_name].values if lat_name != lon_name else None 1133 if lats is not None: 1134 if lons.ndim == 1 and lat_name == "lat": 1135 lons, lats = np.meshgrid(lons, lats) 1136 ps_last = ps_arr[-1] if ps_arr.ndim >= 2 else ps_arr 1137 1138 # For each layer, compare expected fill (PS < layer top bound) 1139 # with actual fill from the output 1140 fill_validation_rows = [] 1141 for k in range(N_VCOARSEN_LAYERS): 1142 p_top = VCOARSEN_PBOUNDS[k] 1143 p_bot = VCOARSEN_PBOUNDS[k + 1] 1144 # Expected fill: surface pressure below the top bound of this layer 1145 expected_fill = (ps_last < p_top).astype(float) 1146 expected_pct = 100.0 * np.sum(expected_fill) / expected_fill.size 1147 1148 vname = f"T_{k}" 1149 arr = get_var(ds_ps, vname) 1150 if arr is not None: 1151 data = arr[-1] if arr.ndim >= 2 else arr 1152 actual_fill = ((np.abs(data) >= 1e10) | ~np.isfinite(data)).astype(float) 1153 actual_pct = 100.0 * np.sum(actual_fill) / actual_fill.size 1154 else: 1155 actual_pct = float('nan') 1156 1157 fill_validation_rows.append({ 1158 "layer": k, 1159 "p_top": p_top / 100, 1160 "p_bot": p_bot / 100, 1161 "expected_pct": expected_pct, 1162 "actual_pct": actual_pct, 1163 }) 1164 1165 # Print fill validation table 1166 print("\n Topographic fill validation (T layers):") 1167 print(f" {'Layer':>5} {'P range (hPa)':>18} {'Expected':>10} {'Actual':>10} {'Match':>6}") 1168 for row in fill_validation_rows: 1169 match = "YES" if abs(row["expected_pct"] - row["actual_pct"]) < 0.1 else "NO" 1170 print(f" {row['layer']:>5} {row['p_top']:>8.1f}-{row['p_bot']:>7.1f}" 1171 f" {row['expected_pct']:>9.2f}% {row['actual_pct']:>9.2f}% {match:>6}") 1172 1173 # 3-panel figure for T_7: PS | T_7 | fill mask overlay 1174 t7_arr = get_var(ds_ps, "T_7") 1175 if t7_arr is not None and HAS_MPL: 1176 t7_last = t7_arr[-1] if t7_arr.ndim >= 2 else t7_arr 1177 expected_fill_7 = (ps_last < VCOARSEN_PBOUNDS[7]).astype(float) 1178 actual_fill_7 = ((np.abs(t7_last) >= 1e10) | ~np.isfinite(t7_last)).astype(float) 1179 1180 panels = [ 1181 (ps_last / 100.0, lons, lats, "Surface Pressure (hPa)"), 1182 (t7_last, lons, lats, "T_7 (907-1013 hPa)"), 1183 (expected_fill_7, lons, lats, "Expected fill (PS < 907 hPa)"), 1184 (actual_fill_7, lons, lats, "Actual fill in T_7"), 1185 ] 1186 1187 # Custom 4-panel figure 1188 fig = plt.figure(figsize=(20, 8)) 1189 cmaps = ["viridis", "RdBu_r", "Reds", "Reds"] 1190 vmins = [400, 200, 0, 0] 1191 vmaxs = [1050, 310, 1, 1] 1192 for idx, (data, lo, la, subtitle) in enumerate(panels): 1193 kw = dict(projection=ccrs.PlateCarree()) if HAS_CARTOPY else {} 1194 ax = fig.add_subplot(2, 2, idx + 1, **kw) 1195 if HAS_CARTOPY: 1196 ax.add_feature(cfeature.COASTLINE, linewidth=0.3) 1197 ax.set_global() 1198 im = _plot_on_ax(ax, data, lo, la, 1199 cmaps[idx], vmins[idx], vmaxs[idx], 1200 HAS_CARTOPY) 1201 plt.colorbar(im, ax=ax, shrink=0.7) 1202 ax.set_title(subtitle, fontsize=10) 1203 fig.suptitle("Topographic Fill Validation: T_7 fill matches PS < 907 hPa", 1204 fontsize=12, y=1.01) 1205 plt.tight_layout() 1206 path = os.path.join(comp_outdir, "eam_topo_fill_validation.png") 1207 fig.savefig(path, dpi=150, bbox_inches="tight") 1208 plt.close(fig) 1209 comp_plots.append(path) 1210 print(f" Wrote eam_topo_fill_validation.png") 1211 1212 # Bar chart: expected vs actual fill per layer 1213 if HAS_MPL and fill_validation_rows: 1214 fig, ax = plt.subplots(figsize=(8, 4)) 1215 layers = [r["layer"] for r in fill_validation_rows] 1216 exp = [r["expected_pct"] for r in fill_validation_rows] 1217 act = [r["actual_pct"] for r in fill_validation_rows] 1218 x = np.arange(len(layers)) 1219 w = 0.35 1220 ax.bar(x - w/2, exp, w, label="Expected (PS < layer top)", color="#4c72b0") 1221 ax.bar(x + w/2, act, w, label="Actual fill in T_k", color="#dd8452") 1222 ax.set_xlabel("Coarsened layer (0=top, 7=surface)") 1223 ax.set_ylabel("Fill fraction (%)") 1224 ax.set_title("Vcoarsen Fill Fraction: Expected from PS vs Actual") 1225 ax.set_xticks(x) 1226 ax.set_xticklabels([f"T_{k}" for k in layers]) 1227 ax.legend() 1228 ax.grid(axis="y", alpha=0.3) 1229 path = os.path.join(comp_outdir, "eam_fill_fraction_validation.png") 1230 fig.savefig(path, dpi=150, bbox_inches="tight") 1231 plt.close(fig) 1232 comp_plots.append(path) 1233 print(f" Wrote eam_fill_fraction_validation.png") 1234 1235 close_ds(ds_ps) 1236 1237 # --- Radiation budget --- 1238 if h0_files: 1239 ds_rad = safe_open(h0_files[0]) 1240 if not has_time_zero(ds_rad) and HAS_XR and isinstance(ds_rad, xr.Dataset): 1241 compute_radiation_budget(ds_rad, comp_outdir, comp_plots) 1242 close_ds(ds_rad) 1243 1244 # --- Time series of key variables across all h0 files --- 1245 if h0_files and HAS_MPL and len(h0_files) >= 1: 1246 ts_vars = ["TS", "PS", "PRECT", "FLUT", "T_7", "STW_7"] 1247 ts_data = {vn: [] for vn in ts_vars} 1248 for fpath in sorted(h0_files): 1249 ds_ts = safe_open(fpath) 1250 if has_time_zero(ds_ts): 1251 close_ds(ds_ts) 1252 continue 1253 for vn in ts_vars: 1254 arr = get_var(ds_ts, vn) 1255 if arr is None: 1256 continue 1257 # iterate over all timesteps in this file 1258 if arr.ndim >= 2: 1259 for tidx in range(arr.shape[0]): 1260 v = valid_data(arr[tidx]) 1261 if v is not None and v.size > 0: 1262 ts_data[vn].append(float(v.mean())) 1263 else: 1264 v = valid_data(arr) 1265 if v is not None and v.size > 0: 1266 ts_data[vn].append(float(v.mean())) 1267 close_ds(ds_ts) 1268 1269 for vn in ts_vars: 1270 vals = ts_data[vn] 1271 if len(vals) >= 2: 1272 time_series(vals, "Timestep index", f"Global Mean {vn}", 1273 vn, comp_outdir, f"eam_timeseries_{vn}.png") 1274 path = os.path.join(comp_outdir, f"eam_timeseries_{vn}.png") 1275 if os.path.exists(path): 1276 comp_plots.append(path) 1277 print(f" Wrote eam_timeseries_{vn}.png") 1278 1279 if comp_plots: 1280 all_plots_by_comp["Comparisons"] = comp_plots 1281 1282 1283 # ───────────────────────────────────────────────────────────────────────────── 1284 # Cross-verification: compare fme_output vs fme_legacy_output 1285 # ───────────────────────────────────────────────────────────────────────────── 1286 1287 def cross_verify(fme_rundir, legacy_rundir, outdir, verbose=False): 1288 """ 1289 Compare online FME output against legacy (raw) output. 1290 1291 Both cases run identical model physics from the same ICs -- the only 1292 difference is what diagnostics get written. Therefore: 1293 1294 1. BFB IDENTITY: any field present in both h0 files must match 1295 bit-for-bit (same model state, same history averaging). 1296 1297 2. ONLINE vs OFFLINE VCOARSEN: the legacy case outputs raw 3D fields 1298 (T, Q, U, V, CLDLIQ, CLDICE, RAINQM at full L80 resolution). 1299 We vcoarsen them offline with the same algorithm / pressure bounds 1300 and compare against the FME case's online T_0..T_7, STW_0..STW_7, 1301 U_0..U_7, V_0..V_7. These should be BFB identical. 1302 1303 3. ONLINE vs OFFLINE DERIVED: legacy raw Q + CLDICE + CLDLIQ + RAINQM 1304 should equal FME STW at every level. After vcoarsening, this tests 1305 derived + vcoarsen together. 1306 1307 Returns a dict of {test_name: list_of_result_dicts} for HTML rendering. 1308 """ 1309 print("\n" + "=" * 70) 1310 print("CROSS-VERIFICATION: fme_output vs fme_legacy_output") 1311 print("=" * 70) 1312 1313 issues = [] 1314 results = [] # list of dicts: {test, field, status, detail} 1315 xv_plots = [] # comparison figures 1316 1317 xv_outdir = os.path.join(outdir, "fme_output", "cross_verify") 1318 os.makedirs(xv_outdir, exist_ok=True) 1319 1320 # -- locate h0 files in both rundirs -------------------------------------- 1321 fme_h0 = find_files(fme_rundir, "*.eam.h0.*.nc") 1322 leg_h0 = find_files(legacy_rundir, "*.eam.h0.*.nc") 1323 1324 if not fme_h0 or not leg_h0: 1325 print(" SKIP EAM: h0 files not found in both rundirs") 1326 print(f" fme: {len(fme_h0)} files, legacy: {len(leg_h0)} files") 1327 return issues 1328 1329 ds_fme = safe_open(fme_h0[0]) 1330 ds_leg = safe_open(leg_h0[0]) 1331 1332 if has_time_zero(ds_fme) or has_time_zero(ds_leg): 1333 print(" SKIP: time dimension has size 0") 1334 close_ds(ds_fme); close_ds(ds_leg) 1335 return issues 1336 1337 # -- 1. FIELD IDENTITY ------------------------------------------------------ 1338 # Detect grid mismatch (FME may be remapped to lat-lon, legacy on native) 1339 fme_dims = get_dims(ds_fme) 1340 leg_dims = get_dims(ds_leg) 1341 grids_match = ("ncol" in fme_dims) == ("ncol" in leg_dims) 1342 1343 if grids_match: 1344 print("\n--- Test 1: BFB Identity (shared fields, same grid) ---") 1345 else: 1346 fme_grid = "lat-lon" if "lat" in fme_dims else "native" 1347 leg_grid = "lat-lon" if "lat" in leg_dims else "native" 1348 print(f"\n--- Test 1: Global-Mean Comparison (FME={fme_grid}, Legacy={leg_grid}) ---") 1349 1350 all_bfb_fields = BFB_INST_FIELDS + BFB_AVG_FIELDS 1351 n_bfb_pass = 0 1352 n_bfb_fail = 0 1353 1354 for var in all_bfb_fields: 1355 fme_data = get_var(ds_fme, var) 1356 leg_data = get_var(ds_leg, var) 1357 if fme_data is None or leg_data is None: 1358 label = "missing-fme" if fme_data is None else "missing-legacy" 1359 results.append({"test": "BFB", "field": var, "status": "SKIP", 1360 "detail": f"{label}"}) 1361 print(f" {var}: SKIP ({label})") 1362 continue 1363 1364 if grids_match and fme_data.shape == leg_data.shape: 1365 # Same grid: point-by-point BFB 1366 diff = np.abs(fme_data.astype(np.float64) - leg_data.astype(np.float64)) 1367 maxdiff = float(np.nanmax(diff)) 1368 status = "BFB" if maxdiff < 1e-12 else "DIFF" 1369 if status == "BFB": 1370 n_bfb_pass += 1 1371 else: 1372 n_bfb_fail += 1 1373 issues.append(f" BFB {var}: max|diff| = {maxdiff:.4g}") 1374 results.append({"test": "BFB", "field": var, "status": status, 1375 "detail": f"max|diff| = {maxdiff:.2e}"}) 1376 tag = "PASS" if status == "BFB" else "FAIL" 1377 print(f" {var}: {tag} max|diff| = {maxdiff:.2e}") 1378 else: 1379 # Different grids: compare area-weighted global means (last timestep) 1380 fme_last = fme_data[-1] if fme_data.ndim >= 2 else fme_data 1381 leg_last = leg_data[-1] if leg_data.ndim >= 2 else leg_data 1382 1383 # Area-weight lat-lon data with cos(lat); native grid is quasi-equal-area 1384 fme_flat = fme_last.ravel().astype(np.float64) if hasattr(fme_last, 'ravel') else fme_last 1385 leg_flat = leg_last.ravel().astype(np.float64) if hasattr(leg_last, 'ravel') else leg_last 1386 1387 fme_lat = get_var(ds_fme, "lat") 1388 if fme_lat is not None and "lat" in fme_dims and fme_last.ndim == 2: 1389 # cos-lat weighting for regular lat-lon grid 1390 coslat = np.cos(np.deg2rad(fme_lat)) 1391 weights = coslat[:, np.newaxis] * np.ones(fme_last.shape[1])[np.newaxis, :] 1392 mask = (np.abs(fme_last) < 1e10) & np.isfinite(fme_last) 1393 fme_mean = float(np.average(fme_last[mask], weights=weights[mask])) if mask.any() else float('nan') 1394 else: 1395 fme_v = valid_data(fme_flat) 1396 fme_mean = float(fme_v.mean()) if fme_v is not None and fme_v.size > 0 else float('nan') 1397 1398 # Native grid (ne30pg2) is quasi-equal-area, unweighted mean is fine 1399 leg_v = valid_data(leg_flat) 1400 leg_mean = float(leg_v.mean()) if leg_v is not None and leg_v.size > 0 else float('nan') 1401 1402 if np.isnan(fme_mean) or np.isnan(leg_mean): 1403 results.append({"test": "GMEAN", "field": var, "status": "SKIP", 1404 "detail": "no valid data"}) 1405 print(f" {var}: SKIP (no valid data)") 1406 continue 1407 1408 rel_diff = abs(fme_mean - leg_mean) / max(abs(leg_mean), 1e-30) 1409 abs_diff = abs(fme_mean - leg_mean) 1410 # For near-zero-mean fields (winds, stresses), relative difference 1411 # is meaningless. Use absolute difference against the larger magnitude. 1412 field_mag = max(abs(fme_mean), abs(leg_mean)) 1413 # Area-weighted means should agree within remap interpolation error (~2%) 1414 # or within 2% of the field magnitude for near-zero fields 1415 if field_mag > 1e-10: 1416 effective_rel = abs_diff / field_mag 1417 else: 1418 effective_rel = 0.0 # both means are essentially zero 1419 status = "PASS" if min(rel_diff, effective_rel) < 0.02 else "DIFF" 1420 if status == "PASS": 1421 n_bfb_pass += 1 1422 else: 1423 n_bfb_fail += 1 1424 issues.append(f" GMEAN {var}: rel_diff = {rel_diff:.4g}") 1425 results.append({"test": "GMEAN", "field": var, "status": status, 1426 "detail": f"fme={fme_mean:.6g}, leg={leg_mean:.6g}, rel={rel_diff:.2e}"}) 1427 print(f" {var}: {status} fme={fme_mean:.6g} leg={leg_mean:.6g} rel={rel_diff:.2e}") 1428 1429 print(f" Summary: {n_bfb_pass} pass, {n_bfb_fail} fail, " 1430 f"{len(all_bfb_fields) - n_bfb_pass - n_bfb_fail} skip") 1431 1432 # -- Native vs Remapped comparison plots ----------------------------------- 1433 if HAS_MPL and not grids_match: 1434 print("\n--- Generating native vs remapped comparison plots ---") 1435 tidx = -1 1436 1437 # Get native grid coordinates (legacy: lat/lon are 1D arrays of ncol) 1438 leg_lat = get_var(ds_leg, "lat") 1439 leg_lon = get_var(ds_leg, "lon") 1440 # Get remapped grid coordinates (FME: lat/lon are 1D coordinate arrays) 1441 fme_lat = get_var(ds_fme, "lat") 1442 fme_lon = get_var(ds_fme, "lon") 1443 1444 if leg_lat is not None and leg_lon is not None and \ 1445 fme_lat is not None and fme_lon is not None: 1446 # Native: 1D arrays for tripcolor 1447 nat_lons = leg_lon if leg_lon.ndim == 1 else leg_lon.ravel() 1448 nat_lats = leg_lat if leg_lat.ndim == 1 else leg_lat.ravel() 1449 # Remapped: meshgrid for pcolormesh 1450 if fme_lat.ndim == 1 and fme_lon.ndim == 1: 1451 rem_lons, rem_lats = np.meshgrid(fme_lon, fme_lat) 1452 else: 1453 rem_lons, rem_lats = fme_lon, fme_lat 1454 1455 plot_specs = [ 1456 # State 1457 ("TS", "RdBu_r", 220, 320, "K"), 1458 ("PS", "viridis", 50000, 105000, "Pa"), 1459 ("ICEFRAC", "Blues", 0, 1, "fraction"), 1460 ("LANDFRAC", "YlGn", 0, 1, "fraction"), 1461 # Surface radiation 1462 ("FSDS", "YlOrRd", 0, 500, "W/m2"), 1463 ("FSUS", "YlOrRd", 0, 300, "W/m2"), 1464 ("FSNS", "YlOrRd", 0, 400, "W/m2"), 1465 ("FLDS", "inferno", 100, 450, "W/m2"), 1466 ("FLUS", "inferno", 150, 550, "W/m2"), 1467 ("FLNS", "inferno", 0, 200, "W/m2"), 1468 # TOA radiation 1469 ("SOLIN", "YlOrRd", 0, 1000, "W/m2"), 1470 ("FSNTOA", "YlOrRd", 0, 500, "W/m2"), 1471 ("FSUTOA", "YlOrRd", 0, 600, "W/m2"), 1472 ("FLUT", "inferno", 100, 320, "W/m2"), 1473 ("FLNT", "inferno", 0, 200, "W/m2"), 1474 # Surface fluxes 1475 ("LHFLX", "YlOrRd", 0, 300, "W/m2"), 1476 ("SHFLX", "RdBu_r", -50, 100, "W/m2"), 1477 ("TAUX", "RdBu_r", -0.5, 0.5, "N/m2"), 1478 ("TAUY", "RdBu_r", -0.3, 0.3, "N/m2"), 1479 ("PRECT", "Blues", 0, 1e-6, "m/s"), 1480 ("DTENDTTW", "RdBu_r", -5e-4, 5e-4, "kg/m2/s"), 1481 ] 1482 for var, cmap, vm, vx, units in plot_specs: 1483 nat_arr = get_var(ds_leg, var) 1484 rem_arr = get_var(ds_fme, var) 1485 if nat_arr is None or rem_arr is None: 1486 continue 1487 nat_data = nat_arr[tidx] if nat_arr.ndim >= 2 else nat_arr 1488 rem_data = rem_arr[tidx] if rem_arr.ndim >= 2 else rem_arr 1489 # Flatten native to 1D for tripcolor 1490 nat_flat = nat_data.ravel() 1491 p = side_by_side_comparison( 1492 nat_flat, nat_lons, nat_lats, f"Native ne30pg2 ({var})", 1493 rem_data, rem_lons, rem_lats, f"Remapped lat-lon ({var})", 1494 f"{var}: Native vs Remapped (last timestep)", 1495 xv_outdir, f"xv_native_vs_remap_{var}.png", 1496 cmap=cmap, vmin=vm, vmax=vx, units=units) 1497 if p: 1498 xv_plots.append(p) 1499 print(f" Wrote xv_native_vs_remap_{var}.png") 1500 1501 # Vcoarsen layers: T_7 and STW_7 (near-surface) 1502 for base, cmap, vm, vx, units in [("T", "RdYlBu_r", 220, 310, "K"), 1503 ("STW", "YlGnBu", 0, 0.025, "kg/kg")]: 1504 vname = f"{base}_7" 1505 fme_vc = get_var(ds_fme, vname) 1506 if fme_vc is not None: 1507 fme_data = fme_vc[tidx] if fme_vc.ndim >= 2 else fme_vc 1508 # No native vcoarsen to compare, just show remapped 1509 p = global_map(fme_data, rem_lons, rem_lats, 1510 f"FME {vname} (remapped, near-surface)", 1511 cmap=cmap, vmin=vm, vmax=vx, 1512 outdir=xv_outdir, fname=f"xv_fme_{vname}_remap.png", 1513 units=units) 1514 if p: 1515 xv_plots.append(p) 1516 print(f" Wrote xv_fme_{vname}_remap.png") 1517 else: 1518 print(" SKIP plots: lat/lon coordinates not found") 1519 1520 # -- 1b. TOPOGRAPHIC FILL VALIDATION ---------------------------------------- 1521 print("\n--- Test 1b: Topographic Fill Validation ---") 1522 # Verify that fill values in coarsened layers correspond exactly to grid cells 1523 # where the surface pressure is below the target layer's top pressure bound. 1524 fme_ps = get_var(ds_fme, "PS") 1525 if fme_ps is not None: 1526 ps_last = fme_ps[-1] if fme_ps.ndim >= 2 else fme_ps 1527 topo_max_err = 0.0 1528 for k in range(N_VCOARSEN_LAYERS): 1529 p_top = VCOARSEN_PBOUNDS[k] 1530 # Expected: fill where PS < layer's top bound (no model levels in range) 1531 expected_pct = 100.0 * np.mean(ps_last.ravel() < p_top) 1532 t_k = get_var(ds_fme, f"T_{k}") 1533 if t_k is None: 1534 continue 1535 t_last = t_k[-1] if t_k.ndim >= 2 else t_k 1536 actual_pct = 100.0 * np.mean( 1537 (np.abs(t_last.ravel()) >= 1e10) | ~np.isfinite(t_last.ravel())) 1538 topo_max_err = max(topo_max_err, abs(expected_pct - actual_pct)) 1539 1540 # Tolerance: remap can shift fill boundaries by ~0.5% of grid cells 1541 topo_status = "PASS" if topo_max_err < 1.0 else "DIFF" 1542 results.append({"test": "TOPO_FILL", "field": "T layers", 1543 "status": topo_status, 1544 "detail": f"max expected-vs-actual fill mismatch: {topo_max_err:.3f}%"}) 1545 print(f" Topographic fill: {topo_status} max mismatch={topo_max_err:.3f}%") 1546 if topo_status != "PASS": 1547 issues.append(f" TOPO_FILL: max fill mismatch {topo_max_err:.3f}%") 1548 else: 1549 print(" SKIP: PS not found in FME output") 1550 1551 # -- 2. ONLINE vs OFFLINE VCOARSEN ---------------------------------------- 1552 print("\n--- Test 2: Online vs Offline Vcoarsen ---") 1553 1554 # Need hybrid coords and PS from legacy file to compute interface pressures 1555 hyai = get_var(ds_leg, "hyai") 1556 hybi = get_var(ds_leg, "hybi") 1557 leg_ps = get_var(ds_leg, "PS") 1558 1559 if hyai is None or hybi is None or leg_ps is None: 1560 print(" SKIP: hyai/hybi/PS not found in legacy file") 1561 else: 1562 # Use the LAST timestep to avoid initialization artifacts 1563 tidx = -1 1564 ps_1t = leg_ps[tidx] # (ncol,) 1565 pint = compute_pint(ps_1t, hyai, hybi) # (ncol, nlev+1) 1566 1567 for fme_base, legacy_sources in FME_TO_LEGACY.items(): 1568 # Build the full-resolution field from legacy 1569 # (for derived fields like STW, sum the components) 1570 raw_3d = None 1571 missing_src = False 1572 for src in legacy_sources: 1573 src_data = get_var(ds_leg, src) 1574 if src_data is None: 1575 print(f" {fme_base}: SKIP (legacy missing {src})") 1576 missing_src = True 1577 break 1578 src_1t = src_data[tidx] # (lev, ncol) or (ncol,) from EAM NetCDF 1579 if src_1t.ndim < 2: 1580 print(f" {fme_base}: SKIP ({src} is not 3D)") 1581 missing_src = True 1582 break 1583 # EAM NetCDF stores 3D as (time, lev, ncol); after time slice: (lev, ncol) 1584 # offline_vcoarsen_avg expects (ncol, lev) — transpose if needed 1585 if src_1t.shape[0] < src_1t.shape[1]: 1586 src_1t = src_1t.T # (lev, ncol) -> (ncol, lev) 1587 if raw_3d is None: 1588 raw_3d = src_1t.astype(np.float64) 1589 else: 1590 raw_3d = raw_3d + src_1t.astype(np.float64) 1591 1592 if missing_src or raw_3d is None: 1593 continue 1594 1595 # Offline vcoarsen 1596 offline_vc = offline_vcoarsen_avg(raw_3d, pint) # (ncol, 8) 1597 1598 # Compare with online vcoarsen from FME output 1599 layer_results = [] 1600 max_err_all = 0.0 1601 max_rel_all = 0.0 1602 for k in range(N_VCOARSEN_LAYERS): 1603 vname = f"{fme_base}_{k}" 1604 fme_arr = get_var(ds_fme, vname) 1605 if fme_arr is None: 1606 layer_results.append(f" {vname}: MISSING in FME output") 1607 continue 1608 1609 fme_1t = fme_arr[tidx].astype(np.float64).ravel() # flatten (lat,lon) or (ncol,) 1610 off_1t = offline_vc[:, k] 1611 1612 # Grids may differ (remapped vs native) — compare area-weighted global means 1613 if fme_1t.size != off_1t.size: 1614 # Area-weight the FME (lat-lon) data 1615 fme_lat = get_var(ds_fme, "lat") 1616 fme_arr_2d = fme_arr[tidx].astype(np.float64) 1617 if fme_lat is not None and fme_arr_2d.ndim == 2: 1618 coslat = np.cos(np.deg2rad(fme_lat)) 1619 wt = coslat[:, np.newaxis] * np.ones(fme_arr_2d.shape[1])[np.newaxis, :] 1620 m = (np.abs(fme_arr_2d) < 1e10) & np.isfinite(fme_arr_2d) 1621 fme_wm = float(np.average(fme_arr_2d[m], weights=wt[m])) if m.any() else float('nan') 1622 else: 1623 fv = valid_data(fme_1t) 1624 fme_wm = float(fv.mean()) if fv is not None else float('nan') 1625 off_v = valid_data(off_1t) 1626 off_wm = float(off_v.mean()) if off_v is not None else float('nan') 1627 if not np.isnan(fme_wm) and not np.isnan(off_wm): 1628 rel = abs(fme_wm - off_wm) / max(abs(off_wm), 1e-30) 1629 max_rel_all = max(max_rel_all, rel) 1630 max_err_all = max(max_err_all, abs(fme_wm - off_wm)) 1631 continue 1632 1633 # Same grid: point-by-point comparison 1634 valid_mask = (np.abs(fme_1t) < 1e10) & (np.abs(off_1t) < 1e10) & \ 1635 np.isfinite(fme_1t) & np.isfinite(off_1t) 1636 1637 if not valid_mask.any(): 1638 layer_results.append(f" {vname}: no valid data") 1639 continue 1640 1641 diff = np.abs(fme_1t[valid_mask] - off_1t[valid_mask]) 1642 maxdiff = float(diff.max()) 1643 scale = np.maximum(np.abs(off_1t[valid_mask]), 1e-30) 1644 maxrel = float((diff / scale).max()) 1645 1646 max_err_all = max(max_err_all, maxdiff) 1647 max_rel_all = max(max_rel_all, maxrel) 1648 1649 # Report — thresholds depend on whether grids match. 1650 # For near-zero fields (U, V, TAUX, TAUY), relative difference is 1651 # meaningless because the global mean crosses zero. Use absolute 1652 # difference instead, benchmarked against a typical scale. 1653 if grids_match: 1654 bfb_thresh, close_thresh = 1e-10, 1e-5 1655 else: 1656 bfb_thresh, close_thresh = 1e-5, 0.02 # remap interpolation ~2% 1657 1658 # Determine a characteristic scale for this field to judge abs diff 1659 off_v = valid_data(offline_vc.ravel()) 1660 field_scale = float(np.percentile(np.abs(off_v), 95)) if off_v is not None and off_v.size > 0 else 1.0 1661 # For near-zero-mean fields, use abs diff / field_scale as the metric 1662 if field_scale > 0: 1663 abs_rel = max_err_all / field_scale 1664 else: 1665 abs_rel = 0.0 1666 1667 # Use the more lenient of relative-mean or absolute-relative 1668 effective_metric = min(max_rel_all, abs_rel) 1669 1670 if effective_metric < bfb_thresh: 1671 status = "BFB" 1672 tag = "PASS" 1673 elif effective_metric < close_thresh: 1674 status = "CLOSE" 1675 tag = "PASS" 1676 else: 1677 status = "DIFF" 1678 tag = "FAIL" 1679 issues.append(f" VCOARSEN {fme_base}: max|rel|={max_rel_all:.2e}, " 1680 f"|diff|/scale={abs_rel:.2e}") 1681 1682 detail = (f"max|diff|={max_err_all:.2e}, max|rel|={max_rel_all:.2e}, " 1683 f"|diff|/p95={abs_rel:.2e}") 1684 results.append({"test": "VCOARSEN", "field": fme_base, 1685 "status": status, "detail": detail}) 1686 print(f" {fme_base}_0..{fme_base}_{N_VCOARSEN_LAYERS-1}: {tag} {detail}") 1687 1688 # Generate difference map for last layer (skip if grids differ) 1689 if HAS_MPL and HAS_XR and grids_match: 1690 last_k = N_VCOARSEN_LAYERS - 1 1691 vname = f"{fme_base}_{last_k}" 1692 fme_arr = get_var(ds_fme, vname) 1693 if fme_arr is not None: 1694 fme_1t = fme_arr[tidx].astype(np.float64) 1695 off_1t = offline_vc[:, last_k] 1696 diff_map = fme_1t - off_1t 1697 1698 # Get coordinates 1699 lon_name = next((d for d in ["lon", "ncol"] if d in get_dims(ds_fme)), None) 1700 lat_var = get_var(ds_fme, "lat") 1701 lon_var = get_var(ds_fme, "lon") 1702 if lat_var is not None and lon_var is not None: 1703 # Only build a meshgrid when lat/lon are true axis arrays 1704 # (different sizes => structured grid). If sizes match 1705 # they are per-column coordinates for an unstructured grid. 1706 if (lat_var.ndim == 1 and lon_var.ndim == 1 1707 and lat_var.size != lon_var.size): 1708 lons, lats = np.meshgrid(lon_var, lat_var) 1709 else: 1710 lons, lats = lon_var.ravel(), lat_var.ravel() 1711 # Side-by-side: online vs offline 1712 _vd_f = valid_data(fme_1t) 1713 vmin_f = float(np.nanpercentile(_vd_f if len(_vd_f) > 0 else [0], 2)) 1714 vmax_f = float(np.nanpercentile(_vd_f if len(_vd_f) > 0 else [0], 98)) 1715 p = side_by_side_comparison( 1716 fme_1t, lons, lats, f"Online {vname}", 1717 off_1t, lons, lats, f"Offline vcoarsen({'+'.join(legacy_sources)})", 1718 f"{fme_base}: Online vs Offline (layer {last_k})", 1719 xv_outdir, f"xv_vcoarsen_{fme_base}_{last_k}.png", 1720 cmap="RdYlBu_r", vmin=vmin_f, vmax=vmax_f) 1721 if p: 1722 xv_plots.append(p) 1723 1724 # -- 3. VCOARSEN LINEARITY: STW_k == Q_k + CLDICE_k + ... ---------------- 1725 print("\n--- Test 3: Vcoarsen Linearity (STW_k = sum of components) ---") 1726 # If we have offline vcoarsen of individual components (Q, CLDICE, CLDLIQ, 1727 # RAINQM), their sum should equal the online STW_k (since vcoarsen is linear). 1728 if hyai is not None and hybi is not None and leg_ps is not None: 1729 tidx = -1 1730 ps_1t = leg_ps[tidx] 1731 pint = compute_pint(ps_1t, hyai, hybi) 1732 1733 # Offline vcoarsen each component 1734 components_vc = {} 1735 all_found = True 1736 for comp in ["Q", "CLDICE", "CLDLIQ", "RAINQM"]: 1737 arr = get_var(ds_leg, comp) 1738 if arr is None or arr.ndim < 3: 1739 all_found = False 1740 break 1741 comp_1t = arr[tidx].astype(np.float64) 1742 if comp_1t.shape[0] < comp_1t.shape[1]: 1743 comp_1t = comp_1t.T # (lev, ncol) -> (ncol, lev) 1744 components_vc[comp] = offline_vcoarsen_avg(comp_1t, pint) 1745 1746 if all_found: 1747 sum_vc = sum(components_vc.values()) # (ncol, 8) 1748 max_lin_err = 0.0 1749 max_lin_rel = 0.0 1750 1751 for k in range(N_VCOARSEN_LAYERS): 1752 stw_k = get_var(ds_fme, f"STW_{k}") 1753 if stw_k is None: 1754 continue 1755 stw_1t = stw_k[tidx].astype(np.float64).ravel() 1756 sum_1t = sum_vc[:, k] 1757 1758 # Different grids: compare global means 1759 if stw_1t.size != sum_1t.size: 1760 sv = valid_data(stw_1t) 1761 ov = valid_data(sum_1t) 1762 if sv is not None and ov is not None: 1763 rel = abs(sv.mean() - ov.mean()) / max(abs(ov.mean()), 1e-30) 1764 max_lin_rel = max(max_lin_rel, rel) 1765 max_lin_err = max(max_lin_err, abs(sv.mean() - ov.mean())) 1766 continue 1767 1768 valid = (np.abs(stw_1t) < 1e10) & np.isfinite(stw_1t) 1769 if not valid.any(): 1770 continue 1771 1772 diff = np.abs(stw_1t[valid] - sum_1t[valid]) 1773 scale = np.maximum(np.abs(sum_1t[valid]), 1e-30) 1774 max_lin_err = max(max_lin_err, float(diff.max())) 1775 max_lin_rel = max(max_lin_rel, float((diff / scale).max())) 1776 1777 if max_lin_rel < 1e-5: 1778 status = "PASS" 1779 else: 1780 status = "FAIL" 1781 issues.append(f" LINEARITY STW: max|rel| = {max_lin_rel:.2e}") 1782 1783 results.append({"test": "LINEARITY", "field": "STW_k", 1784 "status": status, 1785 "detail": f"STW_k vs sum(Q_k+CLDICE_k+CLDLIQ_k+RAINQM_k): " 1786 f"max|rel|={max_lin_rel:.2e}"}) 1787 print(f" STW_k linearity: {status} max|rel|={max_lin_rel:.2e}") 1788 else: 1789 print(" SKIP linearity: component fields not all found in legacy") 1790 1791 close_ds(ds_fme) 1792 close_ds(ds_leg) 1793 1794 # -- 4. TIMING & STORAGE COMPARISON --------------------------------------- 1795 print("\n--- Timing & Storage ---") 1796 1797 # Storage: compare total NetCDF file sizes 1798 def dir_nc_size(d): 1799 total = 0 1800 count = 0 1801 for f in glob.glob(os.path.join(d, "*.nc")): 1802 sz = os.path.getsize(f) 1803 total += sz 1804 count += 1 1805 return total, count 1806 1807 fme_bytes, fme_nfiles = dir_nc_size(fme_rundir) 1808 leg_bytes, leg_nfiles = dir_nc_size(legacy_rundir) 1809 fme_gb = fme_bytes / 1e9 1810 leg_gb = leg_bytes / 1e9 1811 ratio = fme_gb / leg_gb if leg_gb > 0 else float('inf') 1812 print(f" FME output: {fme_nfiles} files, {fme_gb:.2f} GB") 1813 print(f" Legacy output: {leg_nfiles} files, {leg_gb:.2f} GB") 1814 print(f" Storage ratio: {ratio:.2f}x {'(smaller)' if ratio < 1 else '(larger)'}") 1815 1816 results.append({"test": "STORAGE", "field": "total NC files", 1817 "status": "INFO", 1818 "detail": f"FME: {fme_nfiles} files / {fme_gb:.2f} GB, " 1819 f"Legacy: {leg_nfiles} files / {leg_gb:.2f} GB, " 1820 f"ratio: {ratio:.2f}x"}) 1821 1822 # Timing: parse model_timing_stats from both runs 1823 fme_timing = read_timing_summary(fme_rundir) 1824 leg_timing = read_timing_summary(legacy_rundir) 1825 1826 # Try to extract total model time from timing files 1827 def parse_model_time(timing_lines): 1828 """Extract total model run time in seconds from timing summary.""" 1829 if not timing_lines: 1830 return None 1831 for line in timing_lines: 1832 low = line.lower() 1833 # Look for lines with 'total' or 'model' and a number 1834 if ('total' in low or 'model' in low) and ('run' in low or 'time' in low): 1835 parts = line.split() 1836 for p in parts: 1837 try: 1838 val = float(p) 1839 if val > 1: # skip small numbers (likely ratios) 1840 return val 1841 except ValueError: 1842 continue 1843 return None 1844 1845 fme_time = parse_model_time(fme_timing) 1846 leg_time = parse_model_time(leg_timing) 1847 if fme_time and leg_time: 1848 overhead = ((fme_time - leg_time) / leg_time) * 100 1849 print(f" FME model time: {fme_time:.1f} s") 1850 print(f" Legacy model time: {leg_time:.1f} s") 1851 print(f" FME overhead: {overhead:+.1f}%") 1852 results.append({"test": "TIMING", "field": "model run time", 1853 "status": "INFO", 1854 "detail": f"FME: {fme_time:.1f}s, Legacy: {leg_time:.1f}s, " 1855 f"overhead: {overhead:+.1f}%"}) 1856 elif fme_timing or leg_timing: 1857 print(" Timing data found in only one run -- cannot compare") 1858 else: 1859 print(" No timing data found in either run") 1860 1861 # -- Summary -------------------------------------------------------------- 1862 print("\n" + "-" * 50) 1863 n_pass = sum(1 for r in results if r["status"] in ("BFB", "PASS", "CLOSE")) 1864 n_fail = sum(1 for r in results if r["status"] in ("DIFF", "FAIL")) 1865 n_skip = sum(1 for r in results if r["status"] == "SKIP") 1866 n_info = sum(1 for r in results if r["status"] == "INFO") 1867 1868 print(f" CROSS-VERIFY TOTAL: {n_pass} pass, {n_fail} fail, " 1869 f"{n_skip} skip, {n_info} info") 1870 if not issues: 1871 print(" ALL CROSS-VERIFICATION PASSED") 1872 else: 1873 for iss in issues: 1874 print(iss) 1875 1876 return issues, results, xv_plots 1877 1878 1879 def write_cross_verify_html(outdir, results, xv_plots): 1880 """Write the cross-verification section of the HTML dashboard.""" 1881 if not results: 1882 return "" 1883 1884 # Group by test type 1885 tests_by_type = {} 1886 for r in results: 1887 tests_by_type.setdefault(r["test"], []).append(r) 1888 1889 n_total_pass = sum(1 for r in results if r["status"] in ("BFB", "PASS", "CLOSE")) 1890 n_total_fail = sum(1 for r in results if r["status"] in ("DIFF", "FAIL")) 1891 n_total_info = sum(1 for r in results if r["status"] == "INFO") 1892 1893 html = '<h2 id="Cross_Verify">Cross-Verification: FME vs Legacy</h2>\n' 1894 html += '<p>Both cases run identical physics from the same ICs. ' 1895 html += 'FME diagnostics are output-only and do not modify model state. ' 1896 html += f'<strong>{n_total_pass} pass, {n_total_fail} fail, {n_total_info} info</strong></p>\n' 1897 1898 # Tabbed interface for test categories 1899 tab_ids = list(tests_by_type.keys()) 1900 if xv_plots: 1901 tab_ids.append("PLOTS") 1902 1903 html += '<div class="tabs">\n' 1904 for i, tid in enumerate(tab_ids): 1905 labels = { 1906 "BFB": "BFB Identity", "GMEAN": "Global Means", 1907 "TOPO_FILL": "Topo Fill", "VCOARSEN": "Vcoarsen", 1908 "LINEARITY": "Linearity", 1909 "STORAGE": "Storage", "TIMING": "Timing", "PLOTS": "Figures", 1910 } 1911 active = " active" if i == 0 else "" 1912 n_p = sum(1 for r in tests_by_type.get(tid, []) if r["status"] in ("BFB", "PASS", "CLOSE")) 1913 n_t = len(tests_by_type.get(tid, [])) 1914 badge = f' <span class="badge">{n_p}/{n_t}</span>' if tid != "PLOTS" else "" 1915 html += f'<button class="tab{active}" onclick="showTab(\'{tid}\')">{labels.get(tid, tid)}{badge}</button>\n' 1916 html += '</div>\n' 1917 1918 for tid, test_results in tests_by_type.items(): 1919 n_pass = sum(1 for r in test_results if r["status"] in ("BFB", "PASS", "CLOSE")) 1920 n_tot = len(test_results) 1921 status_cls = "pass" if n_pass == n_tot else ("fail" if n_pass < n_tot else "") 1922 1923 labels = { 1924 "BFB": "BFB Identity (shared fields match bit-for-bit)", 1925 "GMEAN": "Area-Weighted Global-Mean Comparison (cos-lat weighted, <2% tolerance)", 1926 "TOPO_FILL": "Topographic Fill Validation (fill matches PS < layer bound)", 1927 "VCOARSEN": "Online vs Offline Vcoarsen (area-weighted global mean comparison)", 1928 "LINEARITY": "Vcoarsen Linearity (STW_k = sum of component_k)", 1929 "STORAGE": "Storage Comparison", "TIMING": "Timing Comparison", 1930 } 1931 vis = "" if tid == tab_ids[0] else ' style="display:none"' 1932 html += f'<div class="tab-content" id="tab_{tid}"{vis}>\n' 1933 html += f'<h3>{labels.get(tid, tid)} ' 1934 html += f'<span class="{status_cls}">({n_pass}/{n_tot})</span></h3>\n' 1935 1936 # Sortable table 1937 html += '<table class="sortable"><thead><tr>' 1938 html += '<th onclick="sortTable(this,0)">Field</th>' 1939 html += '<th onclick="sortTable(this,1)">Status</th>' 1940 html += '<th onclick="sortTable(this,2)">Detail</th></tr></thead><tbody>\n' 1941 1942 for r in test_results: 1943 cls = "pass" if r["status"] in ("BFB", "PASS", "CLOSE") else \ 1944 "fail" if r["status"] in ("DIFF", "FAIL") else "" 1945 html += (f'<tr><td>{r["field"]}</td>' 1946 f'<td class="{cls}">{r["status"]}</td>' 1947 f'<td style="font-family:monospace;font-size:0.85em">{r["detail"]}</td></tr>\n') 1948 html += '</tbody></table>\n' 1949 html += '</div>\n' 1950 1951 # Figures tab 1952 if xv_plots: 1953 vis = "" if "PLOTS" == tab_ids[0] else ' style="display:none"' 1954 html += f'<div class="tab-content" id="tab_PLOTS"{vis}>\n' 1955 html += '<h3>Native vs Remapped Comparison Figures</h3>\n' 1956 html += '<p>Left: native ne30pg2 (tripcolor). Right: remapped Gaussian lat-lon (pcolormesh).</p>\n' 1957 html += '<div class="gallery">\n' 1958 for p in xv_plots: 1959 if p and os.path.exists(p): 1960 rel = os.path.relpath(p, outdir) 1961 html += (f'<div class="fig wide"><a href="{rel}">' 1962 f'<img src="{rel}"/></a>' 1963 f'<span>{os.path.basename(p)}</span></div>\n') 1964 html += '</div>\n</div>\n' 1965 1966 return html 1967 1968 1969 # ----------------------------------------------------------------------------- 1970 # File inventory (EAM only) 1971 # ----------------------------------------------------------------------------- 1972 1973 def collect_file_inventory(rundir): 1974 """Collect EAM file inventory data and print summary. 1975 1976 Returns list of (label, count, file_list) tuples for HTML rendering. 1977 """ 1978 print("\n=== File Inventory ===") 1979 categories = [ 1980 ("EAM h0", "*.eam.h0.*.nc", None), 1981 ] 1982 total = 0 1983 inventory_data = [] 1984 for label, pattern, exclude in categories: 1985 hits = find_files(rundir, pattern, exclude=exclude) 1986 total += len(hits) 1987 status = f"{len(hits)} file(s)" if hits else "NONE" 1988 print(f" {label:38s} {status}") 1989 inventory_data.append((label, len(hits), hits)) 1990 print(f" {'TOTAL':38s} {total} EAM-related file(s)") 1991 return inventory_data 1992 1993 1994 # ----------------------------------------------------------------------------- 1995 # ACE variable coverage 1996 # ----------------------------------------------------------------------------- 1997 1998 def check_variable_coverage(rundir): 1999 """Check which ACE-required variables are present in the FME output. 2000 2001 Returns list of (ace_name, eam_name, category_str, found_bool) tuples. 2002 """ 2003 h0_files = find_files(rundir, "*.eam.h0.*.nc") 2004 if not h0_files: 2005 return [] 2006 2007 ds = safe_open(h0_files[0]) 2008 varnames_in_file = set(get_varnames(ds)) 2009 close_ds(ds) 2010 2011 forcing_set = set(ACE_FORCING_NAMES) 2012 in_set = set(ACE_IN_NAMES) 2013 out_set = set(ACE_OUT_NAMES) 2014 2015 coverage = [] 2016 seen = set() 2017 for ace_name in ACE_FORCING_NAMES + ACE_IN_NAMES + ACE_OUT_NAMES: 2018 if ace_name in seen: 2019 continue 2020 seen.add(ace_name) 2021 eam_name = ace_to_eam(ace_name) 2022 cats = [] 2023 if ace_name in forcing_set: 2024 cats.append("forcing") 2025 if ace_name in in_set: 2026 cats.append("in") 2027 if ace_name in out_set: 2028 cats.append("out") 2029 category = ", ".join(cats) 2030 found = eam_name in varnames_in_file 2031 coverage.append((ace_name, eam_name, category, found)) 2032 2033 return coverage 2034 2035 2036 def generate_yaml_text(): 2037 """Generate YAML-formatted text of the ACE atmosphere variable spec.""" 2038 lines = ["## ATMOSPHERE", "next_step_forcing_names:"] 2039 for name in ACE_FORCING_NAMES: 2040 lines.append(f" - {name}") 2041 lines.append("in_names:") 2042 for name in ACE_IN_NAMES: 2043 lines.append(f" - {name}") 2044 lines.append("out_names:") 2045 for name in ACE_OUT_NAMES: 2046 lines.append(f" - {name}") 2047 return "\n".join(lines) 2048 2049 2050 # ----------------------------------------------------------------------------- 2051 # Reproducibility info 2052 # ----------------------------------------------------------------------------- 2053 2054 def _find_user_nl_eam(rundir): 2055 """Search for user_nl_eam in likely locations relative to rundir.""" 2056 candidates = [ 2057 os.path.join(rundir, "user_nl_eam"), 2058 os.path.join(os.path.dirname(rundir.rstrip("/")), "user_nl_eam"), 2059 ] 2060 for path in candidates: 2061 if os.path.exists(path): 2062 return path 2063 return None 2064 2065 2066 def collect_repro_info(args): 2067 """Collect reproducibility information for the HTML dashboard.""" 2068 import subprocess 2069 2070 info = { 2071 "command": " ".join(sys.argv), 2072 "script_path": os.path.abspath(__file__), 2073 "timestamp": datetime.now().strftime("%Y-%m-%d %H:%M:%S"), 2074 "hostname": os.uname().nodename, 2075 "user": os.environ.get("USER", "unknown"), 2076 "rundir": os.path.abspath(args.rundir), 2077 } 2078 2079 if args.legacy_rundir: 2080 info["legacy_rundir"] = os.path.abspath(args.legacy_rundir) 2081 2082 # Git hash of the script's repo 2083 script_dir = os.path.dirname(info["script_path"]) 2084 try: 2085 result = subprocess.run( 2086 ["git", "-C", script_dir, "rev-parse", "--short", "HEAD"], 2087 capture_output=True, text=True, timeout=5) 2088 if result.returncode == 0: 2089 info["git_hash"] = result.stdout.strip() 2090 result2 = subprocess.run( 2091 ["git", "-C", script_dir, "diff", "--stat", "--", info["script_path"]], 2092 capture_output=True, text=True, timeout=5) 2093 if result2.stdout.strip(): 2094 info["git_dirty"] = True 2095 except Exception: 2096 pass 2097 2098 # Read user_nl_eam from the FME case 2099 nl_path = _find_user_nl_eam(args.rundir) 2100 if nl_path: 2101 try: 2102 with open(nl_path) as f: 2103 info["user_nl_eam"] = f.read() 2104 info["user_nl_eam_path"] = nl_path 2105 except Exception: 2106 pass 2107 2108 # Read user_nl_eam from the legacy case 2109 if args.legacy_rundir: 2110 leg_nl = _find_user_nl_eam(args.legacy_rundir) 2111 if leg_nl: 2112 try: 2113 with open(leg_nl) as f: 2114 info["legacy_user_nl_eam"] = f.read() 2115 info["legacy_user_nl_eam_path"] = leg_nl 2116 except Exception: 2117 pass 2118 2119 # Embed the script source for reproducibility 2120 try: 2121 with open(info["script_path"]) as f: 2122 info["script_source"] = f.read() 2123 except Exception: 2124 pass 2125 2126 return info 2127 2128 2129 # ----------------------------------------------------------------------------- 2130 # HTML helpers for new sections 2131 # ----------------------------------------------------------------------------- 2132 2133 def write_variable_coverage_html(coverage): 2134 """Generate HTML for the ACE Variable Requirements section.""" 2135 if not coverage: 2136 return "" 2137 2138 n_found = sum(1 for _, _, _, f in coverage if f) 2139 n_total = len(coverage) 2140 n_missing = n_total - n_found 2141 pct = 100 * n_found / n_total if n_total else 0 2142 2143 html = '<h2 id="ACE_Variables">ACE Atmosphere Variable Requirements</h2>\n' 2144 2145 # Summary with progress bar 2146 badge_cls = "badge-pass" if n_missing == 0 else "badge-fail" 2147 html += '<div class="card">\n' 2148 html += f'<p style="font-size:1.05em"><strong>{n_found}/{n_total}</strong> variables found ' 2149 if n_missing: 2150 html += f'<span class="badge-fail">{n_missing} missing</span>' 2151 else: 2152 html += '<span class="badge-pass">all present</span>' 2153 html += '</p>\n' 2154 html += f'<div class="progress-bar"><div class="progress-fill" style="width:{pct:.0f}%"></div></div>\n' 2155 html += '</div>\n' 2156 2157 # YAML spec in a collapsible block 2158 yaml_text = generate_yaml_text() 2159 html += '<details><summary>YAML Variable Specification (ACE canonical names)</summary>\n' 2160 html += '<pre class="yaml-block">' 2161 html += yaml_text 2162 html += '</pre>\n</details>\n' 2163 2164 # Name mapping table with filter 2165 html += '<details open><summary>Variable Coverage</summary>\n' 2166 html += '<input class="filter-input" type="text" placeholder="Filter variables..." ' 2167 html += 'oninput="filterTable(this,\'coverage-table\')">\n' 2168 html += '<table class="sortable" id="coverage-table"><thead><tr>' 2169 html += '<th onclick="sortTable(this,0)">ACE Name</th>' 2170 html += '<th onclick="sortTable(this,1)">EAM Name</th>' 2171 html += '<th onclick="sortTable(this,2)">Category</th>' 2172 html += '<th onclick="sortTable(this,3)">Status</th>' 2173 html += '</tr></thead><tbody>\n' 2174 2175 for ace_name, eam_name, category, found in coverage: 2176 cls = "pass" if found else "fail" 2177 status = "FOUND" if found else "MISSING" 2178 html += (f'<tr><td><code>{ace_name}</code></td>' 2179 f'<td><code>{eam_name}</code></td>' 2180 f'<td>{category}</td>' 2181 f'<td class="{cls}">{status}</td></tr>\n') 2182 2183 html += '</tbody></table>\n</details>\n' 2184 return html 2185 2186 2187 def write_repro_html(repro_info): 2188 """Generate HTML for the Reproducibility section.""" 2189 if not repro_info: 2190 return "" 2191 2192 html = '<h2 id="Reproducibility">Reproducibility</h2>\n' 2193 html += '<div class="repro-block"><dl>\n' 2194 2195 html += f'<dt>Command</dt><dd><code>{repro_info["command"]}</code></dd>\n' 2196 html += f'<dt>Script</dt><dd><code>{repro_info["script_path"]}</code>' 2197 if repro_info.get("git_hash"): 2198 dirty = " (dirty)" if repro_info.get("git_dirty") else "" 2199 html += f' — git: <code>{repro_info["git_hash"]}{dirty}</code>' 2200 html += '</dd>\n' 2201 html += f'<dt>Run directory</dt><dd><code>{repro_info["rundir"]}</code></dd>\n' 2202 if repro_info.get("legacy_rundir"): 2203 html += f'<dt>Legacy run directory</dt><dd><code>{repro_info["legacy_rundir"]}</code></dd>\n' 2204 html += f'<dt>Host</dt><dd><code>{repro_info.get("user", "")}@{repro_info.get("hostname", "")}</code></dd>\n' 2205 html += f'<dt>Generated</dt><dd>{repro_info["timestamp"]}</dd>\n' 2206 2207 html += '</dl>\n' 2208 2209 # user_nl_eam contents 2210 if repro_info.get("user_nl_eam"): 2211 html += '<details><summary>user_nl_eam (FME)</summary>\n' 2212 html += f'<pre class="yaml-block">{repro_info["user_nl_eam"]}</pre>\n</details>\n' 2213 2214 if repro_info.get("legacy_user_nl_eam"): 2215 html += '<details><summary>user_nl_eam (Legacy)</summary>\n' 2216 html += f'<pre class="yaml-block">{repro_info["legacy_user_nl_eam"]}</pre>\n</details>\n' 2217 2218 # Full script source with line numbers 2219 if repro_info.get("script_source"): 2220 import html as html_mod 2221 src = repro_info["script_source"] 2222 lines = src.split("\n") 2223 # Build line-numbered source with anchors 2224 numbered = [] 2225 pad = len(str(len(lines))) 2226 for i, line in enumerate(lines, 1): 2227 escaped = html_mod.escape(line) 2228 numbered.append( 2229 f'<span id="L{i}" class="src-line">' 2230 f'<span class="src-ln">{i:>{pad}}</span> {escaped}</span>' 2231 ) 2232 src_html = "\n".join(numbered) 2233 script_name = os.path.basename(repro_info["script_path"]) 2234 html += ( 2235 f'<details><summary>{script_name} ' 2236 f'({len(lines)} lines)</summary>\n' 2237 f'<pre class="src-block">{src_html}</pre>\n' 2238 f'</details>\n' 2239 ) 2240 2241 html += '</div>\n' 2242 return html 2243 2244 2245 # ----------------------------------------------------------------------------- 2246 # Executive summary 2247 # ----------------------------------------------------------------------------- 2248 2249 def build_executive_summary(all_issues, coverage, xv_results=None): 2250 """Compute executive summary metrics from verification results.""" 2251 summary = {} 2252 2253 # Checks passed / total 2254 n_checks_total = 0 2255 n_checks_passed = 0 2256 for comp, issues in all_issues.items(): 2257 n_checks_total += 1 2258 if not issues: 2259 n_checks_passed += 1 2260 summary["n_checks_passed"] = n_checks_passed 2261 summary["n_checks_total"] = n_checks_total 2262 2263 # Variable coverage 2264 if coverage: 2265 n_vars_found = sum(1 for _, _, _, f in coverage if f) 2266 n_vars_total = len(coverage) 2267 else: 2268 n_vars_found = 0 2269 n_vars_total = 0 2270 summary["n_vars_found"] = n_vars_found 2271 summary["n_vars_total"] = n_vars_total 2272 2273 # Storage and timing from cross-verification results 2274 summary["storage_fme_gb"] = None 2275 summary["storage_leg_gb"] = None 2276 summary["storage_ratio"] = None 2277 summary["timing_overhead"] = None 2278 2279 if xv_results: 2280 for r in xv_results: 2281 if r.get("test") == "STORAGE": 2282 detail = r.get("detail", "") 2283 # Parse "FME: N files / X.XX GB, Legacy: M files / Y.YY GB, ratio: Z.ZZx" 2284 import re 2285 m_fme = re.search(r"FME:.*?(\d+\.\d+)\s*GB", detail) 2286 m_leg = re.search(r"Legacy:.*?(\d+\.\d+)\s*GB", detail) 2287 m_rat = re.search(r"ratio:\s*(\d+\.\d+)x", detail) 2288 if m_fme: 2289 summary["storage_fme_gb"] = float(m_fme.group(1)) 2290 if m_leg: 2291 summary["storage_leg_gb"] = float(m_leg.group(1)) 2292 if m_rat: 2293 summary["storage_ratio"] = float(m_rat.group(1)) 2294 elif r.get("test") == "TIMING": 2295 detail = r.get("detail", "") 2296 import re 2297 m_oh = re.search(r"overhead:\s*([+-]?\d+\.?\d*)%", detail) 2298 if m_oh: 2299 summary["timing_overhead"] = float(m_oh.group(1)) 2300 2301 return summary 2302 2303 2304 def write_executive_summary_html(summary): 2305 """Return HTML string with a grid of 4 metric cards.""" 2306 if not summary: 2307 return "" 2308 2309 vars_val = f'{summary["n_vars_found"]}/{summary["n_vars_total"]}' 2310 vars_detail = ("All ACE atmosphere variables found" 2311 if summary["n_vars_found"] == summary["n_vars_total"] 2312 else f'{summary["n_vars_total"] - summary["n_vars_found"]} variables missing') 2313 2314 checks_val = f'{summary["n_checks_passed"]}/{summary["n_checks_total"]}' 2315 checks_detail = ("All component checks passed" 2316 if summary["n_checks_passed"] == summary["n_checks_total"] 2317 else f'{summary["n_checks_total"] - summary["n_checks_passed"]} components with issues') 2318 2319 if summary.get("storage_ratio") is not None: 2320 storage_val = f'{summary["storage_ratio"]:.2f}x' 2321 storage_detail = (f'FME {summary["storage_fme_gb"]:.2f} GB vs ' 2322 f'Legacy {summary["storage_leg_gb"]:.2f} GB') 2323 else: 2324 storage_val = "N/A" 2325 storage_detail = "No cross-verification data" 2326 2327 if summary.get("timing_overhead") is not None: 2328 timing_val = f'{summary["timing_overhead"]:+.1f}%' 2329 timing_detail = "FME overhead vs legacy run" 2330 else: 2331 timing_val = "N/A" 2332 timing_detail = "No timing data available" 2333 2334 html = '<div class="exec-summary">\n' 2335 html += (' <div class="metric-card">\n' 2336 f' <div class="metric-value">{vars_val}</div>\n' 2337 ' <div class="metric-label">Variable Coverage</div>\n' 2338 f' <div class="metric-detail">{vars_detail}</div>\n' 2339 ' </div>\n') 2340 html += (' <div class="metric-card">\n' 2341 f' <div class="metric-value">{checks_val}</div>\n' 2342 ' <div class="metric-label">Checks Passed</div>\n' 2343 f' <div class="metric-detail">{checks_detail}</div>\n' 2344 ' </div>\n') 2345 html += (' <div class="metric-card">\n' 2346 f' <div class="metric-value">{storage_val}</div>\n' 2347 ' <div class="metric-label">Storage Ratio</div>\n' 2348 f' <div class="metric-detail">{storage_detail}</div>\n' 2349 ' </div>\n') 2350 html += (' <div class="metric-card">\n' 2351 f' <div class="metric-value">{timing_val}</div>\n' 2352 ' <div class="metric-label">Timing Overhead</div>\n' 2353 f' <div class="metric-detail">{timing_detail}</div>\n' 2354 ' </div>\n') 2355 html += '</div>\n' 2356 return html 2357 2358 2359 # ----------------------------------------------------------------------------- 2360 # HTML index 2361 # ----------------------------------------------------------------------------- 2362 2363 def write_html_index(outdir, all_plots_by_comp, all_issues, file_inventory_data, 2364 all_fill_reports, timing_summary=None, extra_html="", 2365 coverage=None, repro_info=None, exec_summary=None): 2366 index = os.path.join(outdir, "index.html") 2367 n_issues = sum(len(v) for v in all_issues.values()) 2368 n_pass = sum(1 for v in all_issues.values() if not v) 2369 n_total = len(all_issues) 2370 status = "ALL PASS" if n_issues == 0 else f"{n_issues} issues in {n_total - n_pass}/{n_total} components" 2371 status_color = "#2a7d2a" if n_issues == 0 else "#c33" 2372 2373 # Summary table 2374 summary_rows = "" 2375 for comp, issues in all_issues.items(): 2376 if not issues: 2377 summary_rows += (f'<tr><td>{comp}</td><td class="pass">PASS</td>' 2378 f'<td>0</td></tr>\n') 2379 else: 2380 summary_rows += (f'<tr><td>{comp}</td><td class="fail">FAIL</td>' 2381 f'<td>{len(issues)}</td></tr>\n') 2382 2383 # Detailed issues (collapsible) 2384 detail_rows = "" 2385 for comp, issues in all_issues.items(): 2386 for msg in issues: 2387 detail_rows += f'<tr><td>{comp}</td><td>{msg.strip()}</td></tr>\n' 2388 2389 # File inventory table 2390 inventory_rows = "" 2391 for label, count, flist in file_inventory_data: 2392 status_cls = "pass" if count > 0 else "fail" 2393 inventory_rows += (f'<tr><td>{label}</td>' 2394 f'<td class="{status_cls}">{count}</td></tr>\n') 2395 2396 # Fill/NaN report table 2397 fill_rows = "" 2398 for comp, reports in all_fill_reports.items(): 2399 for r in reports: 2400 pct_cls = "pass" if r["pct"] < 50 else "fail" 2401 fill_rows += (f'<tr><td>{comp}</td><td>{r["name"]}</td>' 2402 f'<td>{r["total"]:,}</td><td>{r["valid"]:,}</td>' 2403 f'<td>{r["fill_nan"]:,}</td>' 2404 f'<td class="{pct_cls}">{r["pct"]:.1f}%</td></tr>\n') 2405 2406 # Group plots by component 2407 fig_sections = "" 2408 for comp, plots in all_plots_by_comp.items(): 2409 if not plots: 2410 continue 2411 anchor = comp.replace(" ", "_").replace("-", "_").replace("(", "").replace(")", "") 2412 fig_sections += f'<h3 id="{anchor}">{comp}</h3>\n<div class="gallery">\n' 2413 # Use wider display for comparison/multi-panel figures 2414 is_comparison = (comp == "Comparisons") 2415 for p in plots: 2416 if p and os.path.exists(p): 2417 rel = os.path.relpath(p, outdir) 2418 fig_cls = "fig wide" if is_comparison else "fig" 2419 fig_sections += (f'<div class="{fig_cls}"><a href="{rel}">' 2420 f'<img src="{rel}"/></a>' 2421 f'<span>{os.path.basename(p)}</span></div>\n') 2422 fig_sections += '</div>\n' 2423 2424 # Navigation 2425 nav_links = "" 2426 nav_items = ["Overview", "Summary", "ACE_Variables", "File_Inventory", "Fill_NaN_Report"] 2427 if extra_html: 2428 nav_items.append("Cross_Verify") 2429 for comp in all_plots_by_comp: 2430 if all_plots_by_comp[comp]: 2431 nav_items.append(comp) 2432 if timing_summary: 2433 nav_items.append("Timing") 2434 if repro_info: 2435 nav_items.append("Reproducibility") 2436 for item in nav_items: 2437 anchor = item.replace(" ", "_").replace("-", "_").replace("(", "").replace(")", "") 2438 display = item.replace("_", " ") 2439 nav_links += f'<a href="#{anchor}">{display}</a>\n' 2440 2441 timing_html = "" 2442 if timing_summary: 2443 timing_rows = "" 2444 for line in timing_summary[:30]: 2445 parts = line.split() 2446 if len(parts) >= 2: 2447 timing_rows += "<tr>" + "".join(f"<td>{p}</td>" for p in parts) + "</tr>\n" 2448 else: 2449 timing_rows += f"<tr><td colspan='6'>{line}</td></tr>\n" 2450 timing_html = f''' 2451 <h2 id="Timing">Performance Timing</h2> 2452 <div class="timing-container"> 2453 <table class="timing-table"> 2454 {timing_rows} 2455 </table> 2456 </div> 2457 ''' 2458 2459 # ACE variable coverage section 2460 coverage_html = write_variable_coverage_html(coverage) if coverage else "" 2461 2462 # Reproducibility section 2463 repro_html = write_repro_html(repro_info) if repro_info else "" 2464 2465 # Executive summary section 2466 exec_summary_html = write_executive_summary_html(exec_summary) if exec_summary else "" 2467 2468 gen_timestamp = datetime.now().strftime("%Y-%m-%d %H:%M:%S") 2469 2470 html = textwrap.dedent(f"""\ 2471 <!DOCTYPE html><html data-theme="light"><head> 2472 <meta charset="utf-8"/> 2473 <meta name="viewport" content="width=device-width, initial-scale=1"/> 2474 <title>EAM FME Output Verification</title> 2475 <style> 2476 :root {{ 2477 --bg: #f0f2f5; --text: #2c3e50; --text-muted: #6c757d; 2478 --card: #ffffff; --card-border: #e2e8f0; 2479 --card-shadow: 0 2px 6px rgba(0,0,0,0.06); 2480 --header-bg: linear-gradient(135deg, #1a2744 0%, #2c3e6b 100%); 2481 --nav-bg: #ffffff; --nav-border: #e2e8f0; 2482 --nav-link-bg: #e8eef4; --nav-link-text: #1a2744; 2483 --th-bg: #1a2744; --th-text: #fff; --th-hover: #2a3d5c; 2484 --accent: #1a2744; --accent-light: #e8eef4; 2485 --pass: #16a34a; --pass-bg: #dcfce7; 2486 --fail: #dc2626; --fail-bg: #fee2e2; 2487 --skip-bg: #fef9c3; --skip: #a16207; 2488 --code-bg: #f1f5f9; --hover-bg: #f8fafc; 2489 --yaml-bg: #1e293b; --yaml-text: #94a3b8; 2490 --stripe: #f8fafc; --border-radius: 8px; 2491 --ring-track: #e2e8f0; --ring-fill: #16a34a; 2492 --toggle-bg: #e2e8f0; --toggle-knob: #1a2744; 2493 }} 2494 [data-theme="dark"] {{ 2495 --bg: #0d1117; --text: #c9d1d9; --text-muted: #8b949e; 2496 --card: #161b22; --card-border: #30363d; 2497 --card-shadow: 0 2px 6px rgba(0,0,0,0.4); 2498 --header-bg: linear-gradient(135deg, #010409 0%, #0d1117 100%); 2499 --nav-bg: #161b22; --nav-border: #30363d; 2500 --nav-link-bg: #21262d; --nav-link-text: #c9d1d9; 2501 --th-bg: #21262d; --th-text: #c9d1d9; --th-hover: #30363d; 2502 --accent: #58a6ff; --accent-light: #1c2d41; 2503 --pass: #3fb950; --pass-bg: #0d2818; 2504 --fail: #f85149; --fail-bg: #3d1418; 2505 --skip-bg: #3d2e00; --skip: #d29922; 2506 --code-bg: #0d1117; --hover-bg: #1c2128; 2507 --yaml-bg: #010409; --yaml-text: #7d8590; 2508 --stripe: #1c2128; --border-radius: 8px; 2509 --ring-track: #30363d; --ring-fill: #3fb950; 2510 --toggle-bg: #30363d; --toggle-knob: #58a6ff; 2511 }} 2512 * {{ box-sizing: border-box; margin: 0; padding: 0; }} 2513 html {{ scroll-behavior: smooth; }} 2514 body {{ font-family: -apple-system, BlinkMacSystemFont, 'Segoe UI', Roboto, 2515 'Helvetica Neue', Arial, sans-serif; 2516 background: var(--bg); color: var(--text); 2517 line-height: 1.6; transition: background 0.3s, color 0.3s; }} 2518 .container {{ max-width: 1400px; margin: 0 auto; padding: 24px 32px; }} 2519 2520 /* Header */ 2521 header {{ background: var(--header-bg); color: #fff; padding: 28px 32px; 2522 display: flex; justify-content: space-between; align-items: center; 2523 flex-wrap: wrap; gap: 12px; }} 2524 header .hdr-left h1 {{ font-size: 1.5em; font-weight: 700; letter-spacing: -0.02em; }} 2525 header .hdr-left .subtitle {{ font-size: 0.85em; opacity: 0.7; margin-top: 2px; }} 2526 header .status-pill {{ display: inline-block; padding: 6px 16px; border-radius: 20px; 2527 font-weight: 700; font-size: 0.95em; letter-spacing: 0.02em; }} 2528 .status-pass {{ background: rgba(22,163,74,0.2); color: #4ade80; }} 2529 .status-fail {{ background: rgba(220,38,38,0.2); color: #fca5a5; }} 2530 .hdr-right {{ display: flex; align-items: center; gap: 16px; }} 2531 2532 /* Theme toggle */ 2533 .theme-toggle {{ background: none; border: 2px solid rgba(255,255,255,0.3); 2534 color: #fff; padding: 6px 14px; border-radius: 20px; 2535 cursor: pointer; font-size: 0.85em; font-weight: 500; 2536 transition: all 0.2s; display: flex; align-items: center; gap: 6px; }} 2537 .theme-toggle:hover {{ border-color: #fff; background: rgba(255,255,255,0.1); }} 2538 .theme-toggle .icon {{ font-size: 1.1em; }} 2539 2540 /* Navigation */ 2541 nav {{ background: var(--nav-bg); padding: 10px 16px; border-radius: var(--border-radius); 2542 box-shadow: var(--card-shadow); border: 1px solid var(--nav-border); 2543 margin: 20px 0; display: flex; flex-wrap: wrap; gap: 6px; 2544 position: sticky; top: 0; z-index: 100; 2545 transition: background 0.3s, border-color 0.3s; }} 2546 nav a {{ color: var(--nav-link-text); text-decoration: none; padding: 6px 14px; 2547 background: var(--nav-link-bg); border-radius: 6px; font-size: 0.82em; 2548 font-weight: 500; transition: all 0.15s; white-space: nowrap; }} 2549 nav a:hover, nav a.active {{ background: var(--accent); color: #fff; }} 2550 2551 /* Section headings */ 2552 h2 {{ color: var(--accent); margin-top: 36px; font-size: 1.25em; font-weight: 700; 2553 padding-bottom: 8px; border-bottom: 2px solid var(--accent); 2554 transition: color 0.3s; }} 2555 h3 {{ color: var(--text); margin-top: 24px; font-size: 1.05em; }} 2556 2557 /* Cards */ 2558 .card {{ background: var(--card); border: 1px solid var(--card-border); 2559 border-radius: var(--border-radius); box-shadow: var(--card-shadow); 2560 padding: 20px; margin-bottom: 16px; transition: all 0.3s; }} 2561 2562 /* Tables */ 2563 table {{ border-collapse: collapse; width: 100%; background: var(--card); 2564 border: 1px solid var(--card-border); border-radius: var(--border-radius); 2565 overflow: hidden; margin-bottom: 16px; transition: all 0.3s; }} 2566 td, th {{ border: 1px solid var(--card-border); padding: 10px 14px; text-align: left; 2567 transition: all 0.3s; }} 2568 th {{ background: var(--th-bg); color: var(--th-text); font-weight: 600; 2569 font-size: 0.88em; text-transform: uppercase; letter-spacing: 0.04em; }} 2570 tbody tr:nth-child(even) {{ background: var(--stripe); }} 2571 tbody tr:hover {{ background: var(--hover-bg); }} 2572 .pass {{ color: var(--pass); font-weight: 700; }} 2573 .fail {{ color: var(--fail); font-weight: 700; }} 2574 td.pass {{ background: var(--pass-bg); }} 2575 td.fail {{ background: var(--fail-bg); }} 2576 code {{ font-family: 'SF Mono', 'Fira Code', 'Cascadia Code', monospace; 2577 font-size: 0.88em; background: var(--code-bg); padding: 2px 6px; 2578 border-radius: 4px; transition: background 0.3s; }} 2579 2580 /* Figure gallery */ 2581 .gallery {{ display: grid; grid-template-columns: repeat(auto-fill, minmax(420px, 1fr)); 2582 gap: 16px; margin-top: 12px; }} 2583 .fig {{ background: var(--card); border: 1px solid var(--card-border); 2584 border-radius: var(--border-radius); overflow: hidden; 2585 transition: all 0.25s; }} 2586 .fig:hover {{ box-shadow: 0 4px 16px rgba(0,0,0,0.12); transform: translateY(-2px); }} 2587 .fig img {{ width: 100%; display: block; cursor: zoom-in; }} 2588 .fig span {{ display: block; font-size: 0.75em; color: var(--text-muted); 2589 padding: 8px 12px; border-top: 1px solid var(--card-border); 2590 font-family: monospace; }} 2591 .fig.wide {{ grid-column: span 2; }} 2592 @media (max-width: 900px) {{ .fig.wide {{ grid-column: span 1; }} }} 2593 2594 /* Details / collapsible */ 2595 details {{ margin: 12px 0; background: var(--card); border-radius: var(--border-radius); 2596 border: 1px solid var(--card-border); transition: all 0.3s; }} 2597 details[open] {{ box-shadow: var(--card-shadow); }} 2598 summary {{ cursor: pointer; font-weight: 600; color: var(--accent); 2599 padding: 12px 16px; list-style: none; display: flex; 2600 align-items: center; gap: 8px; transition: all 0.2s; }} 2601 summary::-webkit-details-marker {{ display: none; }} 2602 summary::before {{ content: '\\25B6'; font-size: 0.7em; transition: transform 0.2s; 2603 display: inline-block; }} 2604 details[open] > summary::before {{ transform: rotate(90deg); }} 2605 summary:hover {{ background: var(--hover-bg); }} 2606 details > :not(summary) {{ padding: 0 16px 16px; }} 2607 2608 /* Timing */ 2609 .timing-container {{ background: var(--card); padding: 16px; 2610 border-radius: var(--border-radius); 2611 border: 1px solid var(--card-border); 2612 overflow-x: auto; max-height: 500px; overflow-y: auto; }} 2613 .timing-table td {{ font-family: monospace; font-size: 0.83em; 2614 padding: 4px 10px; white-space: nowrap; }} 2615 2616 /* YAML / code blocks */ 2617 .yaml-block {{ background: var(--yaml-bg); color: var(--yaml-text); 2618 padding: 16px 20px; border-radius: var(--border-radius); 2619 font-family: 'SF Mono', 'Fira Code', monospace; font-size: 0.83em; 2620 overflow-x: auto; max-height: 400px; overflow-y: auto; 2621 white-space: pre; line-height: 1.5; border: 1px solid var(--card-border); 2622 transition: all 0.3s; }} 2623 2624 /* Repro block */ 2625 .repro-block {{ background: var(--card); padding: 20px 24px; 2626 border-radius: var(--border-radius); 2627 border: 1px solid var(--card-border); 2628 box-shadow: var(--card-shadow); transition: all 0.3s; }} 2629 .repro-block dl {{ margin: 0; display: grid; grid-template-columns: auto 1fr; 2630 gap: 4px 16px; align-items: baseline; }} 2631 .repro-block dt {{ font-weight: 600; color: var(--accent); font-size: 0.88em; 2632 white-space: nowrap; }} 2633 .repro-block dd {{ margin: 0; font-family: monospace; font-size: 0.83em; 2634 color: var(--text-muted); word-break: break-all; }} 2635 2636 /* Progress bar */ 2637 .progress-bar {{ height: 8px; background: var(--ring-track); 2638 border-radius: 4px; overflow: hidden; margin: 8px 0 16px; }} 2639 .progress-fill {{ height: 100%; border-radius: 4px; transition: width 0.6s ease; 2640 background: linear-gradient(90deg, var(--ring-fill), #22d3ee); }} 2641 2642 /* Status badges */ 2643 .badge-pass {{ display: inline-block; padding: 2px 10px; border-radius: 12px; 2644 background: var(--pass-bg); color: var(--pass); 2645 font-weight: 600; font-size: 0.82em; }} 2646 .badge-fail {{ display: inline-block; padding: 2px 10px; border-radius: 12px; 2647 background: var(--fail-bg); color: var(--fail); 2648 font-weight: 600; font-size: 0.82em; }} 2649 .badge-skip {{ display: inline-block; padding: 2px 10px; border-radius: 12px; 2650 background: var(--skip-bg); color: var(--skip); 2651 font-weight: 600; font-size: 0.82em; }} 2652 2653 /* Executive summary */ 2654 .exec-summary {{ display: grid; grid-template-columns: repeat(4, 1fr); 2655 gap: 16px; margin: 20px 0; }} 2656 .metric-card {{ background: var(--card); border: 1px solid var(--card-border); 2657 border-radius: var(--border-radius); padding: 20px 24px; 2658 box-shadow: var(--card-shadow); text-align: center; 2659 transition: all 0.3s; }} 2660 .metric-card:hover {{ box-shadow: 0 4px 16px rgba(0,0,0,0.12); 2661 transform: translateY(-2px); }} 2662 .metric-value {{ font-size: 2em; font-weight: 800; color: var(--accent); 2663 line-height: 1.2; }} 2664 .metric-label {{ font-size: 0.9em; color: var(--text-muted); font-weight: 600; 2665 margin-top: 4px; }} 2666 .metric-detail {{ font-size: 0.78em; color: var(--text-muted); margin-top: 6px; 2667 opacity: 0.8; }} 2668 @media (max-width: 900px) {{ .exec-summary {{ grid-template-columns: repeat(2, 1fr); }} }} 2669 2670 /* Footer */ 2671 footer {{ margin-top: 48px; padding: 20px 32px; 2672 border-top: 1px solid var(--card-border); 2673 color: var(--text-muted); font-size: 0.8em; 2674 background: var(--card); text-align: center; 2675 transition: all 0.3s; }} 2676 2677 /* Lightbox */ 2678 .lightbox {{ display:none; position:fixed; top:0; left:0; width:100%; height:100%; 2679 background:rgba(0,0,0,0.92); z-index:1000; cursor:zoom-out; 2680 align-items:center; justify-content:center; 2681 backdrop-filter: blur(4px); -webkit-backdrop-filter: blur(4px); }} 2682 .lightbox.active {{ display:flex; }} 2683 .lightbox img {{ max-width:94%; max-height:94%; border-radius: 8px; 2684 box-shadow: 0 8px 32px rgba(0,0,0,0.5); 2685 animation: lbFadeIn 0.2s ease; }} 2686 @keyframes lbFadeIn {{ from {{ opacity:0; transform:scale(0.95); }} 2687 to {{ opacity:1; transform:scale(1); }} }} 2688 .lightbox .lb-nav {{ position:absolute; top:50%; transform:translateY(-50%); 2689 color:#fff; font-size:2.5em; cursor:pointer; 2690 padding:20px; opacity:0.5; transition:opacity 0.2s; 2691 user-select:none; }} 2692 .lightbox .lb-nav:hover {{ opacity:1; }} 2693 .lightbox .lb-prev {{ left:10px; }} 2694 .lightbox .lb-next {{ right:10px; }} 2695 .lightbox .lb-close {{ position:absolute; top:16px; right:24px; color:#fff; 2696 font-size:1.8em; cursor:pointer; opacity:0.6; 2697 transition:opacity 0.2s; }} 2698 .lightbox .lb-close:hover {{ opacity:1; }} 2699 2700 /* Tabs */ 2701 .tabs {{ display:flex; flex-wrap:wrap; gap:4px; margin:16px 0 0; }} 2702 .tab {{ padding:8px 18px; border:none; background: var(--nav-link-bg); 2703 color: var(--nav-link-text); cursor:pointer; font-weight:600; 2704 font-size:0.88em; border-radius: 8px 8px 0 0; 2705 transition: all 0.15s; }} 2706 .tab:hover {{ background: var(--hover-bg); }} 2707 .tab.active {{ background: var(--accent); color:#fff; }} 2708 .badge {{ background:rgba(255,255,255,0.2); padding:2px 8px; border-radius:10px; 2709 font-size:0.8em; margin-left:6px; }} 2710 .tab-content {{ background: var(--card); padding:20px; 2711 border-radius:0 var(--border-radius) var(--border-radius) var(--border-radius); 2712 border: 1px solid var(--card-border); 2713 margin-bottom:16px; transition: all 0.3s; }} 2714 2715 /* Sortable headers */ 2716 .sortable th {{ cursor:pointer; user-select:none; position:relative; 2717 padding-right:22px; }} 2718 .sortable th::after {{ content:'\\2195'; position:absolute; right:6px; top:50%; 2719 transform:translateY(-50%); opacity:0.4; font-size:0.85em; }} 2720 .sortable th:hover {{ background: var(--th-hover); }} 2721 2722 /* Filter input */ 2723 .filter-input {{ width: 100%; max-width: 400px; padding: 8px 14px; 2724 border: 1px solid var(--card-border); border-radius: 6px; 2725 background: var(--card); color: var(--text); 2726 font-size: 0.9em; margin-bottom: 12px; 2727 transition: all 0.2s; outline: none; }} 2728 .filter-input:focus {{ border-color: var(--accent); 2729 box-shadow: 0 0 0 3px rgba(88,166,255,0.15); }} 2730 .filter-input::placeholder {{ color: var(--text-muted); }} 2731 2732 /* Embedded source */ 2733 .src-block {{ background: var(--yaml-bg); color: var(--yaml-text); 2734 padding: 0; border-radius: var(--border-radius); 2735 font-family: 'SF Mono', 'Fira Code', 'Cascadia Code', monospace; 2736 font-size: 0.8em; overflow-x: auto; max-height: 600px; 2737 overflow-y: auto; white-space: pre; line-height: 1.55; 2738 border: 1px solid var(--card-border); counter-reset: line; 2739 tab-size: 4; }} 2740 .src-line {{ display: block; padding: 0 16px 0 0; }} 2741 .src-line:hover {{ background: rgba(88,166,255,0.08); }} 2742 .src-line:target {{ background: rgba(88,166,255,0.18); }} 2743 .src-ln {{ display: inline-block; width: 4.5em; text-align: right; 2744 color: rgba(139,148,158,0.4); padding: 0 12px 0 12px; 2745 user-select: none; border-right: 1px solid var(--card-border); 2746 margin-right: 12px; }} 2747 2748 /* Animations */ 2749 @keyframes fadeIn {{ from {{ opacity:0; transform:translateY(8px); }} 2750 to {{ opacity:1; transform:translateY(0); }} }} 2751 .card, table, details {{ animation: fadeIn 0.3s ease; }} 2752 </style> 2753 <script> 2754 /* Theme toggle */ 2755 (function() {{ 2756 var saved = localStorage.getItem('eam-theme'); 2757 if (saved) document.documentElement.setAttribute('data-theme', saved); 2758 else if (window.matchMedia && window.matchMedia('(prefers-color-scheme: dark)').matches) 2759 document.documentElement.setAttribute('data-theme', 'dark'); 2760 }})(); 2761 function toggleTheme() {{ 2762 var html = document.documentElement; 2763 var next = html.getAttribute('data-theme') === 'dark' ? 'light' : 'dark'; 2764 html.setAttribute('data-theme', next); 2765 localStorage.setItem('eam-theme', next); 2766 var btn = document.querySelector('.theme-toggle .icon'); 2767 if (btn) btn.textContent = next === 'dark' ? '\\u2600' : '\\u263E'; 2768 }} 2769 2770 /* Tabs */ 2771 function showTab(id) {{ 2772 document.querySelectorAll('.tab-content').forEach(function(el) {{ el.style.display='none'; }}); 2773 document.querySelectorAll('.tab').forEach(function(el) {{ el.classList.remove('active'); }}); 2774 var tab = document.getElementById('tab_' + id); 2775 if (tab) tab.style.display = ''; 2776 document.querySelectorAll('.tab').forEach(function(btn) {{ 2777 if (btn.getAttribute('onclick') && btn.getAttribute('onclick').indexOf(id) >= 0) 2778 btn.classList.add('active'); 2779 }}); 2780 }} 2781 2782 /* Sortable tables */ 2783 function sortTable(th, col) {{ 2784 var table = th.closest('table'); 2785 var tbody = table.querySelector('tbody'); 2786 if (!tbody) return; 2787 var rows = Array.from(tbody.querySelectorAll('tr')); 2788 var asc = th.dataset.sort !== 'asc'; 2789 th.dataset.sort = asc ? 'asc' : 'desc'; 2790 // Reset other headers 2791 th.closest('tr').querySelectorAll('th').forEach(function(h) {{ 2792 if (h !== th) delete h.dataset.sort; 2793 }}); 2794 rows.sort(function(a, b) {{ 2795 var va = a.cells[col].textContent.trim(); 2796 var vb = b.cells[col].textContent.trim(); 2797 var na = parseFloat(va), nb = parseFloat(vb); 2798 if (!isNaN(na) && !isNaN(nb)) return asc ? na - nb : nb - na; 2799 return asc ? va.localeCompare(vb) : vb.localeCompare(va); 2800 }}); 2801 rows.forEach(function(r) {{ tbody.appendChild(r); }}); 2802 }} 2803 2804 /* Table filter */ 2805 function filterTable(input, tableId) {{ 2806 var filter = input.value.toLowerCase(); 2807 var table = document.getElementById(tableId); 2808 if (!table) return; 2809 var rows = table.querySelectorAll('tbody tr'); 2810 rows.forEach(function(row) {{ 2811 var text = row.textContent.toLowerCase(); 2812 row.style.display = text.indexOf(filter) > -1 ? '' : 'none'; 2813 }}); 2814 }} 2815 2816 document.addEventListener('DOMContentLoaded', function() {{ 2817 /* Theme button label */ 2818 var theme = document.documentElement.getAttribute('data-theme'); 2819 var icon = document.querySelector('.theme-toggle .icon'); 2820 if (icon) icon.textContent = theme === 'dark' ? '\\u2600' : '\\u263E'; 2821 2822 /* Lightbox with navigation */ 2823 var lb = document.createElement('div'); 2824 lb.className = 'lightbox'; 2825 lb.innerHTML = '<span class="lb-close">\\u00D7</span>' 2826 + '<span class="lb-nav lb-prev">\\u276E</span>' 2827 + '<img/>' 2828 + '<span class="lb-nav lb-next">\\u276F</span>'; 2829 document.body.appendChild(lb); 2830 var lbImg = lb.querySelector('img'); 2831 var allFigs = []; 2832 var curIdx = 0; 2833 2834 function openLB(idx) {{ 2835 curIdx = idx; 2836 lbImg.src = allFigs[idx]; 2837 lb.classList.add('active'); 2838 }} 2839 function closeLB() {{ lb.classList.remove('active'); }} 2840 function navLB(dir) {{ 2841 curIdx = (curIdx + dir + allFigs.length) % allFigs.length; 2842 lbImg.src = allFigs[curIdx]; 2843 }} 2844 2845 document.querySelectorAll('.fig a').forEach(function(a, i) {{ 2846 allFigs.push(a.href); 2847 a.addEventListener('click', function(e) {{ 2848 e.preventDefault(); 2849 openLB(i); 2850 }}); 2851 }}); 2852 2853 lb.querySelector('.lb-close').addEventListener('click', closeLB); 2854 lb.querySelector('.lb-prev').addEventListener('click', function(e) {{ e.stopPropagation(); navLB(-1); }}); 2855 lb.querySelector('.lb-next').addEventListener('click', function(e) {{ e.stopPropagation(); navLB(1); }}); 2856 lb.addEventListener('click', function(e) {{ if (e.target === lb) closeLB(); }}); 2857 document.addEventListener('keydown', function(e) {{ 2858 if (!lb.classList.contains('active')) return; 2859 if (e.key === 'Escape') closeLB(); 2860 if (e.key === 'ArrowLeft') navLB(-1); 2861 if (e.key === 'ArrowRight') navLB(1); 2862 }}); 2863 2864 /* Active nav highlighting on scroll */ 2865 var sections = document.querySelectorAll('h2[id]'); 2866 var navLinks = document.querySelectorAll('nav a'); 2867 if (sections.length && navLinks.length) {{ 2868 var observer = new IntersectionObserver(function(entries) {{ 2869 entries.forEach(function(entry) {{ 2870 if (entry.isIntersecting) {{ 2871 var id = entry.target.getAttribute('id'); 2872 navLinks.forEach(function(link) {{ 2873 link.classList.toggle('active', link.getAttribute('href') === '#' + id); 2874 }}); 2875 }} 2876 }}); 2877 }}, {{ rootMargin: '-20% 0px -70% 0px' }}); 2878 sections.forEach(function(s) {{ observer.observe(s); }}); 2879 }} 2880 }}); 2881 </script> 2882 </head><body> 2883 <header> 2884 <div class="hdr-left"> 2885 <h1>EAM FME Online Output Verification</h1> 2886 <div class="subtitle">E3SM Atmosphere Model — Full Model Emulation Diagnostics</div> 2887 </div> 2888 <div class="hdr-right"> 2889 <span class="status-pill {"status-pass" if n_issues == 0 else "status-fail"}">{status}</span> 2890 <button class="theme-toggle" onclick="toggleTheme()"><span class="icon"></span> Theme</button> 2891 </div> 2892 </header> 2893 <div class="container"> 2894 2895 <nav>{nav_links}</nav> 2896 2897 <h2 id="Overview">Overview</h2> 2898 {exec_summary_html} 2899 2900 <h2 id="Summary">Component Summary</h2> 2901 <table> 2902 <tr><th>Component</th><th>Status</th><th>Issues</th></tr> 2903 {summary_rows} 2904 </table> 2905 2906 {"<details><summary>Show all " + str(n_issues) + " issues</summary><table><tr><th>Component</th><th>Issue</th></tr>" + detail_rows + "</table></details>" if detail_rows else ""} 2907 2908 {coverage_html} 2909 2910 <h2 id="File_Inventory">File Inventory</h2> 2911 <table> 2912 <tr><th>Category</th><th>File Count</th></tr> 2913 {inventory_rows} 2914 </table> 2915 2916 <h2 id="Fill_NaN_Report">Fill / NaN Report</h2> 2917 <details><summary>Variable-level fill fraction details</summary> 2918 <table> 2919 <tr><th>Component</th><th>Variable</th><th>Total Cells</th><th>Valid</th><th>Fill/NaN</th><th>Fill %</th></tr> 2920 {fill_rows if fill_rows else '<tr><td colspan="6">No data collected.</td></tr>'} 2921 </table> 2922 </details> 2923 2924 {extra_html} 2925 2926 <h2>Diagnostic Figures</h2> 2927 {fig_sections if fig_sections else '<p>No figures generated.</p>'} 2928 2929 {timing_html} 2930 2931 {repro_html} 2932 </div> 2933 <footer> 2934 <strong>verify_eam.py</strong> — EAM FME Online Output Verification for E3SM 2935 — Generated {gen_timestamp} 2936 </footer> 2937 </body></html> 2938 """) 2939 with open(index, "w") as fh: 2940 fh.write(html) 2941 print(f"\nIndex written: {index}") 2942 return index 2943 2944 2945 # ----------------------------------------------------------------------------- 2946 # Main 2947 # ----------------------------------------------------------------------------- 2948 2949 def main(): 2950 parser = argparse.ArgumentParser( 2951 description="Verify EAM FME output (vcoarsen, derived fields) and " 2952 "produce diagnostic figures with optional cross-verification.") 2953 parser.add_argument("--rundir", required=True, 2954 help="CIME RUNDIR for the FME case (fme_output testmod)") 2955 parser.add_argument("--outdir", required=True, 2956 help="Output directory for figures and HTML index") 2957 parser.add_argument("--verbose", "-v", action="store_true") 2958 parser.add_argument("--legacy-rundir", 2959 help="CIME RUNDIR for the legacy case (fme_legacy_output testmod) " 2960 "for cross-verification") 2961 args = parser.parse_args() 2962 2963 if not os.path.isdir(args.rundir): 2964 sys.exit(f"ERROR: rundir not found: {args.rundir}") 2965 2966 os.makedirs(args.outdir, exist_ok=True) 2967 print(f"EAM FME Verification | rundir: {args.rundir}") 2968 if args.legacy_rundir: 2969 print(f"Legacy RUNDIR : {args.legacy_rundir}") 2970 print(f"Output directory : {args.outdir}") 2971 print("=" * 70) 2972 2973 if not HAS_MPL: 2974 print("WARNING: matplotlib not available -- no figures will be produced") 2975 if not HAS_CARTOPY: 2976 print("WARNING: cartopy not available -- falling back to scatter maps") 2977 if not (HAS_XR or HAS_NC4): 2978 sys.exit("ERROR: need xarray or netCDF4") 2979 2980 # File inventory 2981 file_inventory_data = collect_file_inventory(args.rundir) 2982 2983 all_issues = {} 2984 all_plots_by_comp = {} 2985 all_fill_reports = {} 2986 2987 # All figures go under fme_output/ subdir so fme_output.html can reference them 2988 fig_root = os.path.join(args.outdir, "fme_output") 2989 os.makedirs(fig_root, exist_ok=True) 2990 2991 def run_check(name, func, subdir, *a): 2992 comp_outdir = os.path.join(fig_root, subdir) 2993 os.makedirs(comp_outdir, exist_ok=True) 2994 result = func(*a[:1], comp_outdir, *a[1:]) 2995 comp_issues, plots, fill_reports = result 2996 all_issues[name] = comp_issues 2997 all_plots_by_comp[name] = plots 2998 all_fill_reports[name] = fill_reports 2999 3000 run_check("EAM", check_eam, "eam", 3001 args.rundir, args.verbose) 3002 3003 # EAM comparison figures (multi-panel layer views, zonal means) 3004 generate_comparison_figures(args.rundir, fig_root, all_plots_by_comp) 3005 3006 # Timing summary 3007 timing_summary = read_timing_summary(args.rundir) 3008 if timing_summary: 3009 print("\n=== Performance Timing ===") 3010 for line in timing_summary[:10]: 3011 print(f" {line}") 3012 if len(timing_summary) > 10: 3013 print(f" ... ({len(timing_summary) - 10} more lines in HTML report)") 3014 3015 # ACE variable coverage 3016 print("\n=== ACE Variable Coverage ===") 3017 coverage = check_variable_coverage(args.rundir) 3018 if coverage: 3019 n_found = sum(1 for _, _, _, f in coverage if f) 3020 n_missing = len(coverage) - n_found 3021 print(f" {n_found}/{len(coverage)} ACE variables found in output") 3022 if n_missing: 3023 for ace_name, eam_name, _, found in coverage: 3024 if not found: 3025 print(f" MISSING: {ace_name} (EAM: {eam_name})") 3026 3027 # Reproducibility info 3028 repro_info = collect_repro_info(args) 3029 3030 # Cross-verification against legacy testmod output 3031 xv_html = "" 3032 xv_results = None 3033 if args.legacy_rundir: 3034 if not os.path.isdir(args.legacy_rundir): 3035 print(f"WARNING: legacy-rundir not found: {args.legacy_rundir}") 3036 else: 3037 xv_issues, xv_results, xv_plots = cross_verify( 3038 args.rundir, args.legacy_rundir, args.outdir, args.verbose) 3039 all_issues["Cross-Verify"] = xv_issues 3040 if xv_plots: 3041 all_plots_by_comp["Cross-Verify"] = xv_plots 3042 xv_html = write_cross_verify_html(args.outdir, xv_results, xv_plots) 3043 3044 # Build executive summary 3045 exec_summary = build_executive_summary(all_issues, coverage, 3046 xv_results=xv_results) 3047 3048 write_html_index(args.outdir, all_plots_by_comp, all_issues, 3049 file_inventory_data, all_fill_reports, 3050 timing_summary=timing_summary, 3051 extra_html=xv_html, 3052 coverage=coverage, 3053 repro_info=repro_info, 3054 exec_summary=exec_summary) 3055 3056 n_total = sum(len(v) for v in all_issues.values()) 3057 print("\n" + "=" * 70) 3058 print(f"RESULT: {'ALL CHECKS PASSED' if n_total == 0 else f'{n_total} ISSUES FOUND'}") 3059 sys.exit(0 if n_total == 0 else 1) 3060 3061 3062 if __name__ == "__main__": 3063 main() 3064